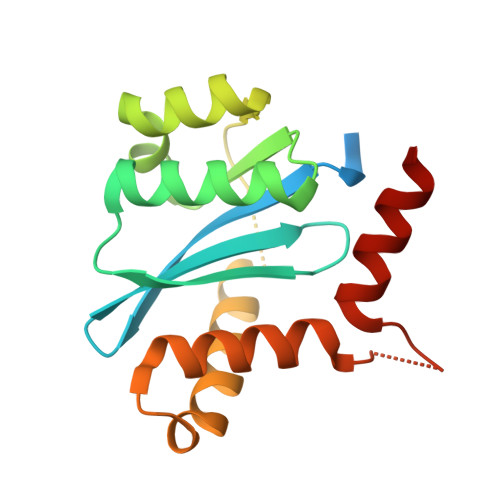
5,6,7,8-Tetrahydro-1,6-naphthyridine Derivatives as Potent HIV-1-Integrase-Allosteric-Site Inhibitors.
Peese, K.M., Allard, C.W., Connolly, T., Johnson, B.L., Li, C., Patel, M., Sorensen, M.E., Walker, M.A., Meanwell, N.A., McAuliffe, B., Minassian, B., Krystal, M., Parker, D.D., Lewis, H.A., Kish, K., Zhang, P., Nolte, R.T., Simmermacher, J., Jenkins, S., Cianci, C., Naidu, B.N.(2019) J Med Chem 62: 1348-1361
- PubMed: 30609350 Search on PubMed
- DOI: https://doi.org/10.1021/acs.jmedchem.8b01473
- Primary Citation Related Structures:
6NCJ - PubMed Abstract:
A series of 5,6,7,8-tetrahydro-1,6-naphthyridine derivatives targeting the allosteric lens-epithelium-derived-growth-factor-p75 (LEDGF/p75)-binding site on HIV-1 integrase, an attractive target for antiviral chemotherapy, was prepared and screened for activity against HIV-1 infection in cell culture. Small molecules that bind within the LEDGF/p75-binding site promote aberrant multimerization of the integrase enzyme and are of significant interest as HIV-1-replication inhibitors. Structure-activity-relationship studies and rat pharmacokinetic studies of lead compounds are presented.
- Protein Cellular and Structural Sciences , GlaxoSmithKline , 1250 South Collegeville Rd. , Collegeville , Pennsylvania 19426 , United States.
Organizational Affiliation: