Conformational Plasticity in the HIV-1 Fusion Peptide Facilitates Recognition by Broadly Neutralizing Antibodies.
Yuan, M., Cottrell, C.A., Ozorowski, G., van Gils, M.J., Kumar, S., Wu, N.C., Sarkar, A., Torres, J.L., de Val, N., Copps, J., Moore, J.P., Sanders, R.W., Ward, A.B., Wilson, I.A.(2019) Cell Host Microbe 25: 873
- PubMed: 31194940 Search on PubMedSearch on PubMed Central
- DOI: https://doi.org/10.1016/j.chom.2019.04.011
- Primary Citation Related Structures:
6NC2, 6NC3, 6NCP - PubMed Abstract:
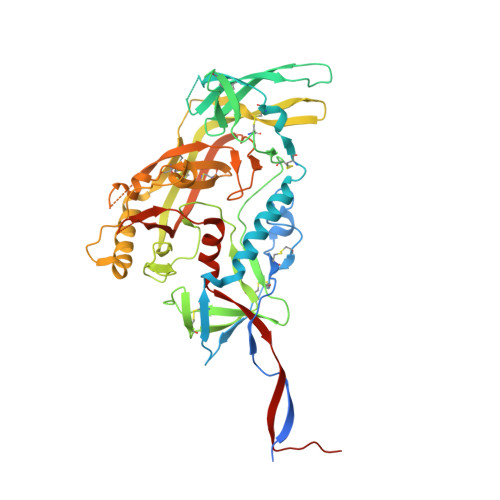
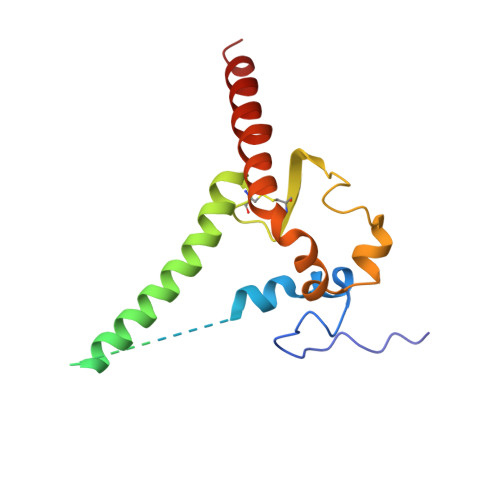
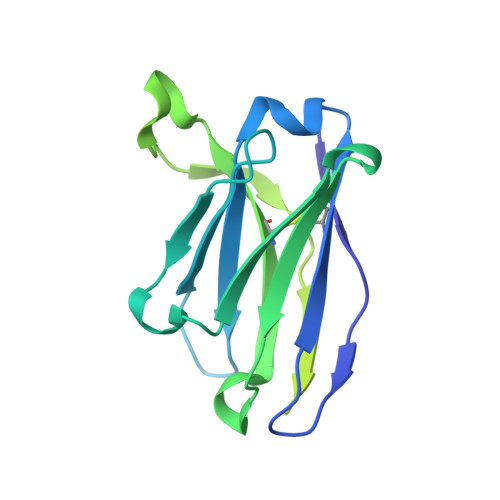
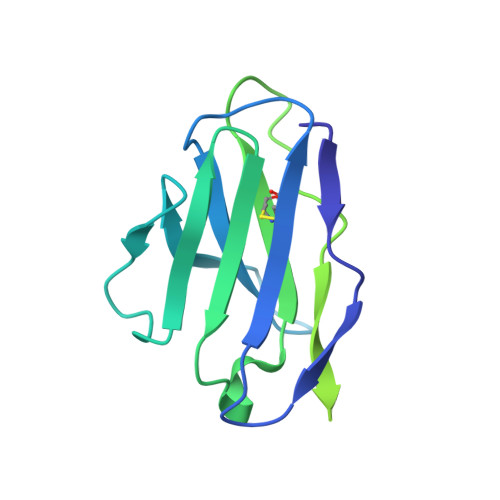
The fusion peptide (FP) of HIV-1 envelope glycoprotein (Env) is essential for mediating viral entry. Detection of broadly neutralizing antibodies (bnAbs) that interact with the FP has revealed it as a site of vulnerability. We delineate X-ray and cryo-electron microscopy (cryo-EM) structures of bnAb ACS202, from an HIV-infected elite neutralizer, with an FP and with a soluble Env trimer (AMC011 SOSIP.v4.2) derived from the same patient. We show that ACS202 CDRH3 forms a "β strand" interaction with the exposed hydrophobic FP and recognizes a continuous region of gp120, including a conserved N-linked glycan at N88. A cryo-EM structure of another previously identified bnAb VRC34.01 with AMC011 SOSIP.v4.2 shows that it also penetrates through glycans to target the FP. We further demonstrate that the FP can twist and present different conformations for recognition by bnAbs, which enables approach to Env from diverse angles. The variable recognition of FP by bnAbs thus provides insights for vaccine design.
- Department of Integrative Structural and Computational Biology, The Scripps Research Institute, La Jolla, CA 92037, USA.
Organizational Affiliation: