Discovery of Adamantane Carboxamides as Ebola Virus Cell Entry and Glycoprotein Inhibitors.
Plewe, M.B., Sokolova, N.V., Gantla, V.R., Brown, E.R., Naik, S., Fetsko, A., Lorimer, D.D., Dranow, D.M., Smutney, H., Bullen, J., Sidhu, R., Master, A., Wang, J., Kallel, E.A., Zhang, L., Kalveram, B., Freiberg, A.N., Henkel, G., McCormack, K.(2020) ACS Med Chem Lett 11: 1160-1167
- PubMed: 32550996 Search on PubMedSearch on PubMed Central
- DOI: https://doi.org/10.1021/acsmedchemlett.0c00025
- Primary Citation Related Structures:
6NAE - PubMed Abstract:
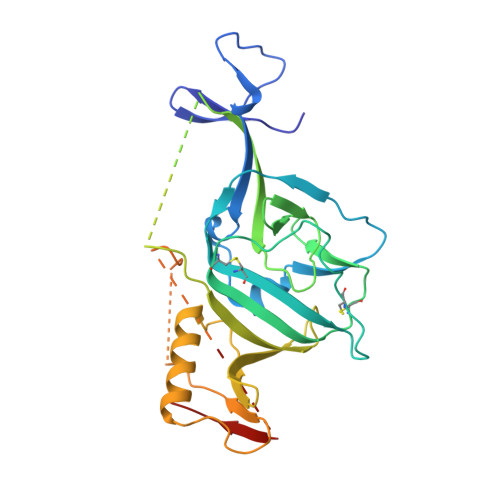
We identified and explored the structure-activity-relationship (SAR) of an adamantane carboxamide chemical series of Ebola virus (EBOV) inhibitors. Selected analogs exhibited half-maximal inhibitory concentrations (EC 50 values) of ∼10-15 nM in vesicular stomatitis virus (VSV) pseudotyped EBOV (pEBOV) infectivity assays, low hundred nanomolar EC 50 activity against wild type EBOV, aqueous solubility >20 mg/mL, and attractive metabolic stability in human and nonhuman liver microsomes. X-ray cocrystallographic characterizations of a lead compound with the EBOV glycoprotein (GP) established the EBOV GP as a target for direct compound inhibitory activity and further provided relevant structural models that may assist in identifying optimized therapeutic candidates.
- Arisan Therapeutics, 11189 Sorrento Valley Road, Suite 104, San Diego, California 92121, United States.
Organizational Affiliation: