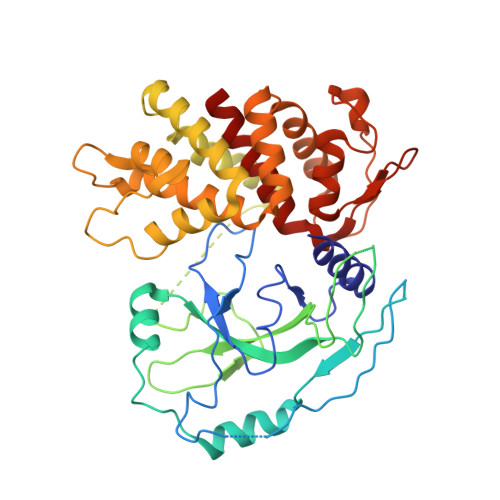
Crystal Structures of the Full-Length Murine and Human Gasdermin D Reveal Mechanisms of Autoinhibition, Lipid Binding, and Oligomerization.
Liu, Z., Wang, C., Yang, J., Zhou, B., Yang, R., Ramachandran, R., Abbott, D.W., Xiao, T.S.(2019) Immunity 51: 43
- PubMed: 31097341 Search on PubMedSearch on PubMed Central
- DOI: https://doi.org/10.1016/j.immuni.2019.04.017
- Primary Citation Related Structures:
6N9N, 6N9O - PubMed Abstract:
Gasdermin D (GSDMD) is an effector molecule for pyroptosis downstream of canonical and noncanonical inflammasome signaling pathways. Cleavage of GSDMD by inflammatory caspases triggers the oligomerization and lipid binding by its N-terminal domain, which assembles membrane pores, whereas its C-terminal domain binds the N-terminal domain to inhibit pyroptosis. Despite recent progress in our understanding of the structure and function of the murine gasdermin A3 (mGSDMA3), the molecular mechanisms of GSDMD activation and regulation remain poorly characterized. Here, we report the crystal structures of the full-length murine and human GSDMDs, which reveal the architecture of the GSDMD N-terminal domains and demonstrate distinct and common features of autoinhibition among gasdermin family members utilizing their β1-β2 loops. Disruption of the intramolecular domain interface enhanced pyroptosis, whereas mutations at the predicted lipid-binding or oligomerization surface reduced cytolysis. Our study provides a framework for understanding the autoinhibition, lipid binding, and oligomerization of GSDMD by using overlapping interfaces.
- Department of Pathology, Case Western Reserve University, Cleveland, OH 44106 USA.
Organizational Affiliation: