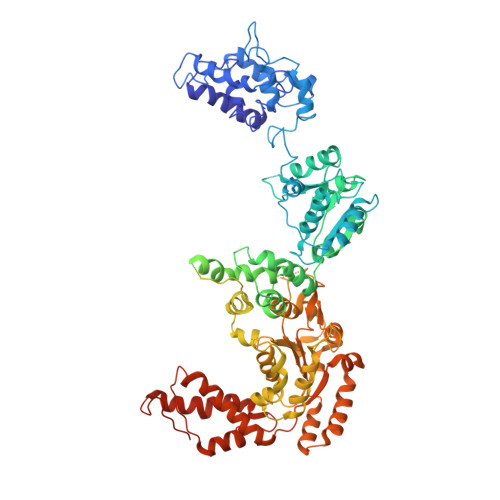
Cryo-EM Structures of the Hsp104 Protein Disaggregase Captured in the ATP Conformation.
Lee, S., Roh, S.H., Lee, J., Sung, N., Liu, J., Tsai, F.T.F.(2019) Cell Rep 26: 29-36.e3
- PubMed: 30605683 Search on PubMedSearch on PubMed Central
- DOI: https://doi.org/10.1016/j.celrep.2018.12.037
- Primary Citation Related Structures:
6N8T, 6N8V, 6N8Z - PubMed Abstract:
Hsp104 is a ring-forming, ATP-driven molecular machine that recovers functional protein from both stress-denatured and amyloid-forming aggregates. Although Hsp104 shares a common architecture with Clp/Hsp100 protein unfoldases, different and seemingly conflicting 3D structures have been reported. Examining the structure of Hsp104 poses considerable challenges because Hsp104 readily hydrolyzes ATP, whereas ATP analogs can be slowly turned over and are often contaminated with other nucleotide species. Here, we present the single-particle electron cryo-microscopy (cryo-EM) structures of a catalytically inactive Hsp104 variant (Hsp104 DWB ) in the ATP-bound state determined between 7.7 Å and 9.3 Å resolution. Surprisingly, we observe that the Hsp104 DWB hexamer adopts distinct ring conformations (closed, extended, and open) despite being in the same nucleotide state. The latter underscores the structural plasticity of Hsp104 in solution, with different conformations stabilized by nucleotide binding. Our findings suggest that, in addition to ATP hydrolysis-driven conformational changes, Hsp104 uses stochastic motions to translocate unfolded polypeptides.
- Verna and Marrs McLean Department of Biochemistry and Molecular Biology, Baylor College of Medicine, Houston, TX 77030, USA. Electronic address: slee@bcm.edu.
Organizational Affiliation: