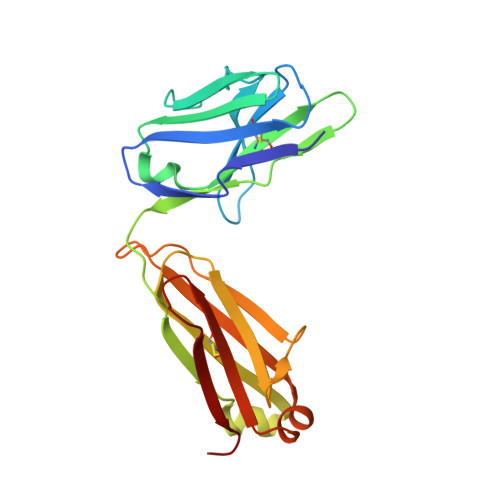
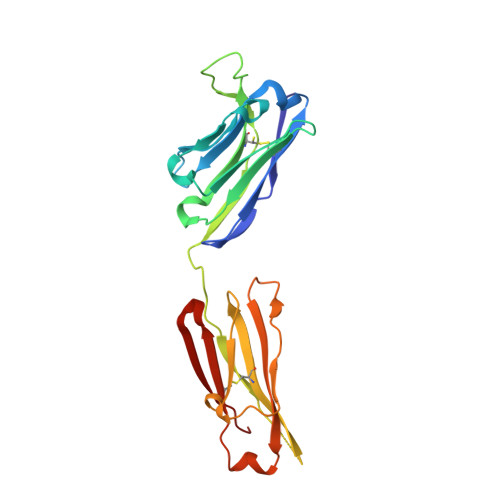
Antibody domain exchange is an immunological solution to carbohydrate cluster recognition.
Calarese, D.A., Scanlan, C.N., Zwick, M.B., Deechongkit, S., Mimura, Y., Kunert, R., Zhu, P., Wormald, M.R., Stanfield, R.L., Roux, K.H., Kelly, J.W., Rudd, P.M., Dwek, R.A., Katinger, H., Burton, D.R., Wilson, I.A.(2003) Science 300: 2065-2071
- PubMed: 12829775 Search on PubMed
- DOI: https://doi.org/10.1126/science.1083182
- Primary Citation Related Structures:
6N2X, 6N32, 6N35 - PubMed Abstract:
Human antibody 2G12 neutralizes a broad range of human immunodeficiency virus type 1 (HIV-1) isolates by binding an unusually dense cluster of carbohydrate moieties on the "silent" face of the gp120 envelope glycoprotein. Crystal structures of Fab 2G12 and its complexes with the disaccharide Manalpha1-2Man and with the oligosaccharide Man9GlcNAc2 revealed that two Fabs assemble into an interlocked VH domain-swapped dimer. Further biochemical, biophysical, and mutagenesis data strongly support a Fab-dimerized antibody as the prevalent form that recognizes gp120. The extraordinary configuration of this antibody provides an extended surface, with newly described binding sites, for multivalent interaction with a conserved cluster of oligomannose type sugars on the surface of gp120. The unique interdigitation of Fab domains within an antibody uncovers a previously unappreciated mechanism for high-affinity recognition of carbohydrate or other repeating epitopes on cell or microbial surfaces.
- Department of Molecular Biology, The Scripps Research Institute, 10550 North Torrey Pines Road, La Jolla, CA 92037, USA.
Organizational Affiliation: