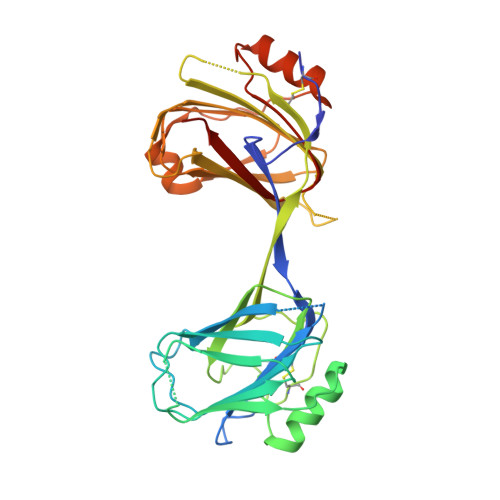
Structural Adaptation in Its Orphan Domain Engenders Betaglycan with an Alternate Mode of Growth Factor Binding Relative to Endoglin.
Kim, S.K., Whitley, M.J., Krzysiak, T.C., Hinck, C.S., Taylor, A.B., Zwieb, C., Byeon, C.H., Zhou, X., Mendoza, V., Lopez-Casillas, F., Furey, W., Hinck, A.P.(2019) Structure 27: 1427-1442.e4
- PubMed: 31327662 Search on PubMedSearch on PubMed Central
- DOI: https://doi.org/10.1016/j.str.2019.06.010
- Primary Citation Related Structures:
6MZN, 6MZP - PubMed Abstract:
Betaglycan (BG) and endoglin (ENG), homologous co-receptors of the TGF-β family, potentiate the signaling activity of TGF-β2 and inhibin A, and BMP-9 and BMP-10, respectively. BG exists as monomer and forms 1:1 growth factor (GF) complexes, while ENG exists as a dimer and forms 2:1 GF complexes. Herein, the structure of the BG orphan domain (BG O ) reveals an insertion that blocks the region that the endoglin orphan domain (ENG O ) uses to bind BMP-9, preventing it from binding in the same manner. Using binding studies with domain-deleted forms of TGF-β and BG O , as well as small-angle X-ray scattering data, BG O is shown to bind its cognate GF in an entirely different manner compared with ENG O . The alternative interfaces likely engender BG and ENG with the ability to selectively bind and target their cognate GFs in a unique temporal-spatial manner, without interfering with one another or other TGF-β family GFs.
- Department of Structural Biology, University of Pittsburgh School of Medicine, Biomedical Science Tower 3, Room 2051, 3501 Fifth Avenue, Pittsburgh, PA 15260, USA; Department of Biochemistry and Structural Biology, University of Texas Health Science Center at San Antonio, San Antonio, TX 78229-3900, USA.
Organizational Affiliation: