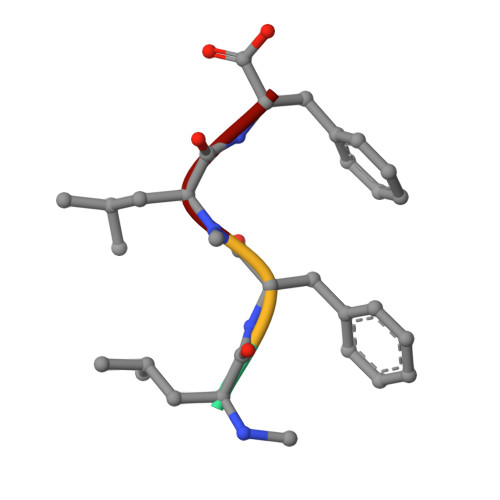
Investigations of the key macrolactamisation step in the synthesis of cyclic tetrapeptide pseudoxylallemycin A.
Cameron, A.J., Squire, C.J., Gerenton, A., Stubbing, L.A., Harris, P.W.R., Brimble, M.A.(2019) Org Biomol Chem 17: 3902-3913
- PubMed: 30941386 Search on PubMed
- DOI: https://doi.org/10.1039/c9ob00227h
- Primary Citation Related Structures:
6MVZ, 6MW0, 6MW1, 6MW2 - PubMed Abstract:
The total synthesis and structural confirmation of naturally occurring all l-cyclic tetrapeptide pseudoxylallemycin A is reported. X-ray crystallography revealed that the linear precursor adopted an all-trans (ttt) extended linear conformation, while its cyclic derivative adopts a trans,cis,trans,cis (tctc) conformation. Two kinetically favoured cyclic conformers prone to hydrolysis initially formed rapidly during cyclisation, with subsequent conversion to the thermodynamically stable tctc macrocycle taking place slowly. We postulate the initial unstable cyclic product undergoes an unprecedented nucleophilic ring opening with either the T3P or PyAOP by-products to give the linear ttt structure as a reactivated species and through a series of equilibria is slowly consumed by cyclisation to the thermodynamic product pseudoxylallemycin A. Consumption of the reactivated species by formation of pseudoxylallemycin A requires a trans-cis isomerism to occur and necessitates moderately increased reaction temperatures. Cyclisation with T3P was found to provide the greatest stereoretention. Synthesis and X-ray crystallography of the C-terminal epimer demonstrated its cyclisation to be kinetically favoured and to proceed without epimerisation despite also bearing an all-trans backbone.
- School of Chemical Sciences, The University of Auckland, 23 Symonds St, Auckland 1010, New Zealand. m.brimble@auckland.ac.nz.
Organizational Affiliation: