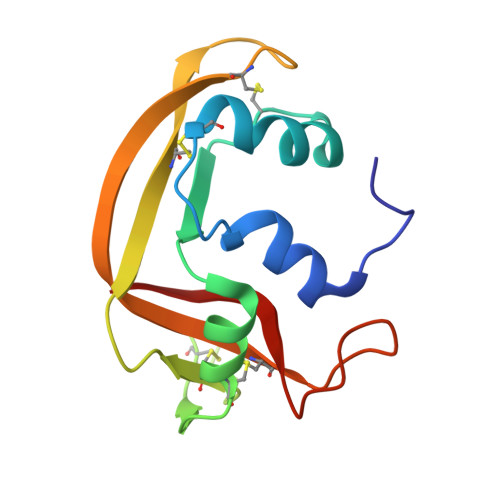
Insights into Structural and Dynamical Changes Experienced by Human RNase 6 upon Ligand Binding.
Narayanan, C., Bernard, D.N., Letourneau, M., Gagnon, J., Gagne, D., Bafna, K., Calmettes, C., Couture, J.F., Agarwal, P.K., Doucet, N.(2020) Biochemistry 59: 755-765
- PubMed: 31909602 Search on PubMedSearch on PubMed Central
- DOI: https://doi.org/10.1021/acs.biochem.9b00888
- Primary Citation Related Structures:
6MV6, 6MV7 - PubMed Abstract:
Ribonuclease 6 (RNase 6) is one of eight catalytically active human pancreatic-type RNases that belong to a superfamily of rapidly evolving enzymes. Like some of its human homologues, RNase 6 exhibits host defense properties such as antiviral and antibacterial activities. Recently solved crystal structures of this enzyme in its nucleotide-free form show the conservation of the prototypical kidney-shaped fold preserved among vertebrate RNases, in addition to revealing the presence of a unique secondary active site. In this study, we determine the structural and conformational properties experienced by RNase 6 upon binding to substrate and product analogues. We present the first crystal structures of RNase 6 bound to a nucleotide ligand (adenosine 5'-monophosphate), in addition to RNase 6 bound to phosphate ions. While the enzyme preserves B 2 subsite ligand preferences, our results show a lack of typical B 2 subsite interactions normally observed in homologous ligand-bound RNases. A comparison of the dynamical properties of RNase 6 in its apo-, substrate-, and product-bound states highlight the unique dynamical properties experienced on time scales ranging from nano- to milliseconds. Overall, our results confirm the specific evolutionary adaptation of RNase 6 relative to its unique catalytic and biological activities.
- Centre Armand-Frappier Santé Biotechnologie , Institut National de la Recherche Scientifique (INRS), Université du Québec , 531 Boulevard des Prairies , Laval , Quebec City H7V 1B7 , Canada.
Organizational Affiliation: