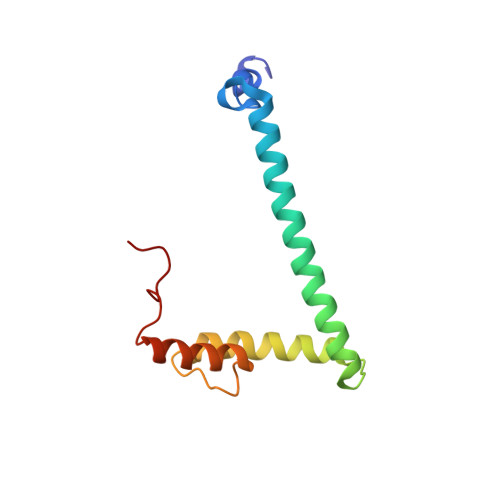
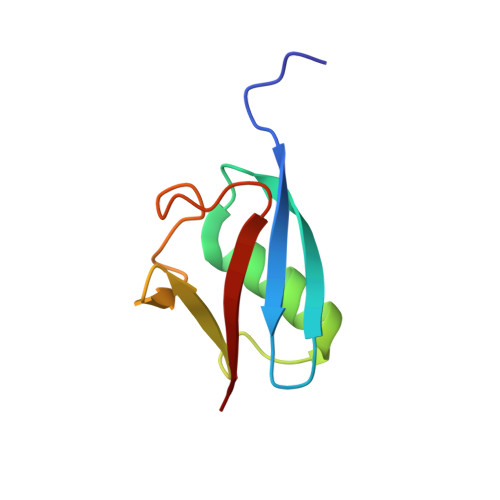
Structure of hRpn10 Bound to UBQLN2 UBL Illustrates Basis for Complementarity between Shuttle Factors and Substrates at the Proteasome.
Chen, X., Ebelle, D.L., Wright, B.J., Sridharan, V., Hooper, E., Walters, K.J.(2019) J Mol Biology 431: 939-955
- PubMed: 30664872 Search on PubMedSearch on PubMed Central
- DOI: https://doi.org/10.1016/j.jmb.2019.01.021
- Primary Citation Related Structures:
6MUN - PubMed Abstract:
The 26S proteasome is a highly complex 2.5-MDa molecular machine responsible for regulated protein degradation. Proteasome substrates are typically marked by ubiquitination for recognition at receptor sites contributed by Rpn1/S2/PSMD2, Rpn10/S5a, and Rpn13/Adrm1. Each receptor site can bind substrates directly by engaging conjugated ubiquitin chains or indirectly by binding to shuttle factors Rad23/HR23, Dsk2/PLIC/UBQLN, or Ddi1, which contain a ubiquitin-like domain (UBL) that adopts the ubiquitin fold. Previous structural studies have defined how each of the proteasome receptor sites binds to ubiquitin chains as well as some of the interactions that occur with the shuttle factors. Here, we define how hRpn10 binds to the UBQLN2 UBL domain, solving the structure of this complex by NMR, and determine affinities for each UIM region by a titration experiment. UBQLN2 UBL exhibits 25-fold stronger affinity for the N-terminal UIM-1 over UIM-2 of hRpn10. Moreover, we discover that UBQLN2 UBL is fine-tuned for the hRpn10 UIM-1 site over the UIM-2 site by taking advantage of the additional contacts made available through the longer UIM-1 helix. We also test hRpn10 versatility for the various ubiquitin chains to find less specificity for any particular linkage type compared to hRpn1 and hRpn13, as expected from the flexible linker region that connects the two UIMs; nonetheless, hRpn10 does exhibit some preference for K48 and K11 linkages. Altogether, these results provide new insights into the highly complex and complementary roles of the proteasome receptor sites and shuttle factors.
- Protein Processing Section, Structural Biophysics Laboratory, Center for Cancer Research, National Cancer Institute, Frederick, MD 21702, USA.
Organizational Affiliation: