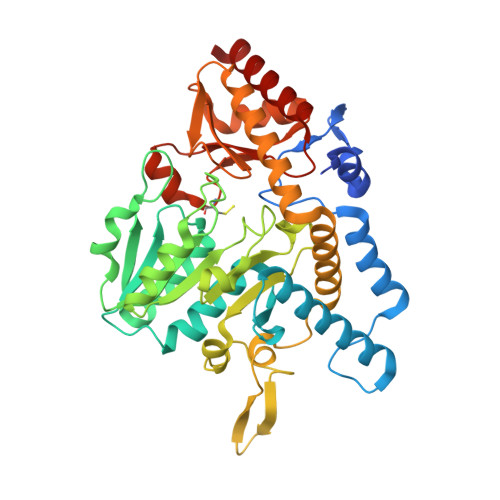
Structural Evidence for Dimer-Interface-Driven Regulation of the Type II Cysteine Desulfurase, SufS.
Dunkle, J.A., Bruno, M.R., Outten, F.W., Frantom, P.A.(2019) Biochemistry 58: 687-696
- PubMed: 30571100 Search on PubMedSearch on PubMed Central
- DOI: https://doi.org/10.1021/acs.biochem.8b01122
- Primary Citation Related Structures:
6MR2, 6MR6, 6MRE, 6MRH, 6MRI - PubMed Abstract:
SufS is a type II cysteine desulfurase and acts as the initial step in the Suf Fe-S cluster assembly pathway. In Escherichia coli, this pathway is utilized under conditions of oxidative stress and is resistant to reactive oxygen species. Mechanistically, this means SufS must shift between protecting a covalent persulfide intermediate and making it available for transfer to the next protein partner in the pathway, SufE. Here, we report five X-ray crystal structures of SufS including a new structure of SufS containing an inward-facing persulfide intermediate on C364. Additional structures of SufS variants with substitutions at the dimer interface show changes in dimer geometry and suggest a conserved β-hairpin structure plays a role in mediating interactions with SufE. These new structures, along with previous HDX-MS and biochemical data, identify an interaction network capable of communication between active-sites of the SufS dimer coordinating the shift between desulfurase and transpersulfurase activities.
- Department of Chemistry & Biochemistry , The University of Alabama , Tuscaloosa , Alabama 35487 , United States.
Organizational Affiliation: