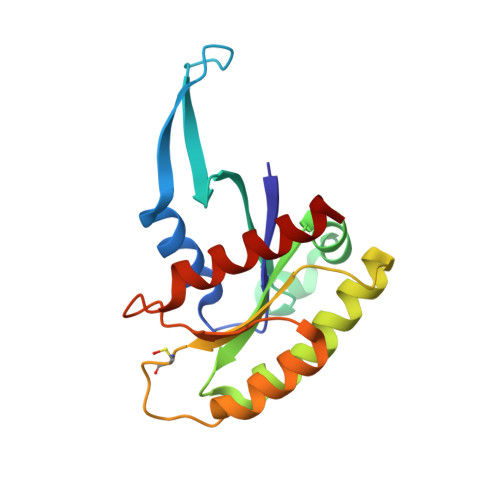
Structural basis of the atypical activation mechanism of KRASV14I.
Bera, A.K., Lu, J., Wales, T.E., Gondi, S., Gurbani, D., Nelson, A., Engen, J.R., Westover, K.D.(2019) J Biological Chem 294: 13964-13972
- PubMed: 31341022 Search on PubMedSearch on PubMed Central
- DOI: https://doi.org/10.1074/jbc.RA119.009131
- Primary Citation Related Structures:
6MQG, 6MQN - PubMed Abstract:
RAS regulation and signaling are largely accomplished by direct protein-protein interactions, making RAS protein dynamics a critical determinant of RAS function. Here, we report a crystal structure of GDP-bound KRAS V14I , a mutated KRAS variant associated with the developmental RASopathy disorder Noonan syndrome (NS), at 1.5-1.6 Å resolution. The structure is notable for revealing a marked extension of switch 1 away from the G-domain and nucleotide-binding site of the KRAS protein. We found that this extension is associated with a loss of the magnesium ion and a tilt in the position of the guanine base because of the additional carbon introduced by the isoleucine substitution. Hydrogen-deuterium exchange MS analysis confirmed that this conformation occurs in solution, but also disclosed a difference in kinetics when compared with KRAS A146T , another RAS mutant that displays a nearly identical conformation in previously reported crystal structures. This conformational change contributed to a high rate of guanine nucleotide-exchange factor (GEF)-dependent and -independent nucleotide exchange and to an increase in affinity for SOS Ras/Rac GEF 1 (SOS1), which appears to be the major mode of activation for this RAS variant. These results highlight a mechanistic connection between KRAS A146T and KRAS V14I that may have implications for the regulation of these variants and for the development of therapeutic strategies to manage KRAS variant-associated disorders.
- Departments of Biochemistry and Radiation Oncology, The University of Texas Southwestern Medical Center at Dallas, Dallas, Texas 75390.
Organizational Affiliation: