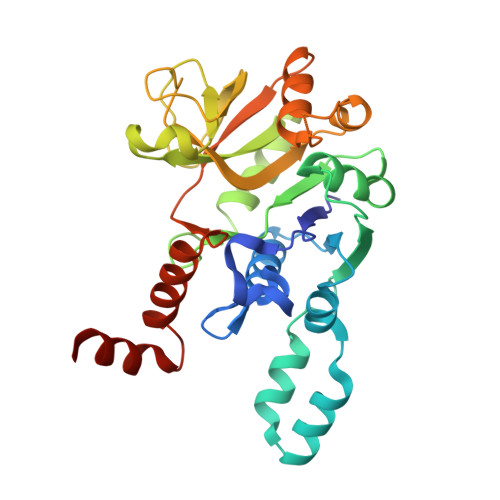
Crystal structure of UDP-glucose pyrophosphorylase from Yersinia pestis, a potential therapeutic target against plague.
Gibbs, M.E., Lountos, G.T., Gumpena, R., Waugh, D.S.(2019) Acta Crystallogr F Struct Biol Commun 75: 608-615
- PubMed: 31475928
- DOI: https://doi.org/10.1107/S2053230X19011154
- Primary Citation Related Structures:
6MNU - PubMed Abstract:
Yersinia pestis, the causative agent of bubonic plague, is one of the most lethal pathogens in recorded human history. Today, the concern is the possible misuse of Y. pestis as an agent in bioweapons and bioterrorism. Current therapies for the treatment of plague include the use of a small number of antibiotics, but clinical cases of antibiotic resistance have been reported in some areas of the world. Therefore, the discovery of new drugs is required to combat potential Y. pestis infection. Here, the crystal structure of the Y. pestis UDP-glucose pyrophosphorylase (UGP), a metabolic enzyme implicated in the survival of Y. pestis in mouse macrophages, is described at 2.17 Å resolution. The structure provides a foundation that may enable the rational design of inhibitors and open new avenues for the development of antiplague therapeutics.
- Macromolecular Crystallography Laboratory, Center for Cancer Research, National Cancer Institute at Frederick, Frederick, MD 21702, USA.
Organizational Affiliation: