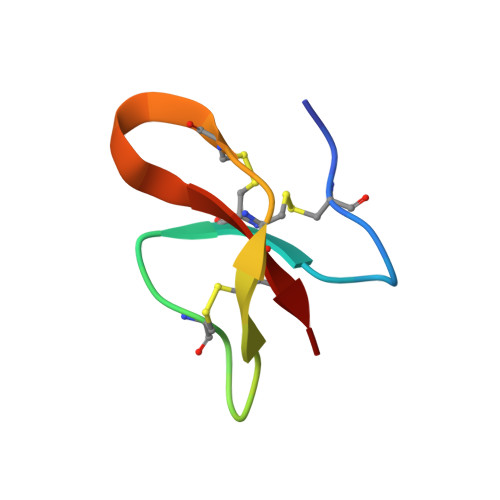
Antimicrobial activity and structure of a consensus human beta-defensin and its comparison to a novel putative hBD10.
Rodriguez, A., Pedersen, M.O., Villegas, E., Rivas-Santiago, B., Villegas-Moreno, J., Amero, C., Norton, R.S., Corzo, G.(2020) Proteins 88: 175-186
- PubMed: 31325337
- DOI: https://doi.org/10.1002/prot.25785
- Primary Citation Related Structures:
6MJV - PubMed Abstract:
The spread of multidrug resistant bacteria owing to the intensive use of antibiotics is challenging current antibiotic therapies, and making the discovery and evaluation of new antimicrobial agents a high priority. The evaluation of novel peptide sequences of predicted antimicrobial peptides from different sources is valuable approach to identify alternative antibiotic leads. Two strategies were pursued in this study to evaluate novel antimicrobial peptides from the human β-defensin family (hBD). In the first, a 32-residue peptide was designed based on the alignment of all available hBD primary structures, while in the second a putative 35-residue peptide, hBD10, was mined from the gene DEFB110. Both hBDconsensus and hBD10 were chemically synthesized, folded and purified. They showed antimicrobial activity against Escherichia coli, Staphylococcus aureus, and Mycobacterium tuberculosis, but were not hemolytic on human red blood cells. The NMR-based solution structure of hBDconsensus revealed that it adopts a classical β-defensin fold and disulfide connectivities. Even though the mass spectrum of hBD10 confirmed the formation of three disulfide bonds, it showed limited dispersion in 1 H NMR spectra and structural studies were not pursued. The evaluation of different β-defensin structures may identify new antimicrobial agents effective against multidrug-resistant bacterial strains.
- Centro de Investigación en Biotecnología, Universidad Autónoma del Estado de Morelos, Cuernavaca, Mexico.
Organizational Affiliation: