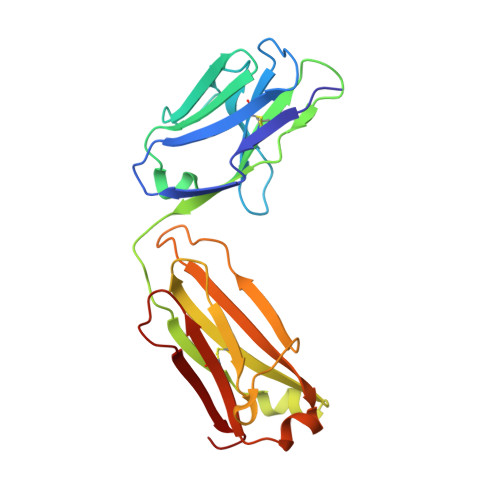
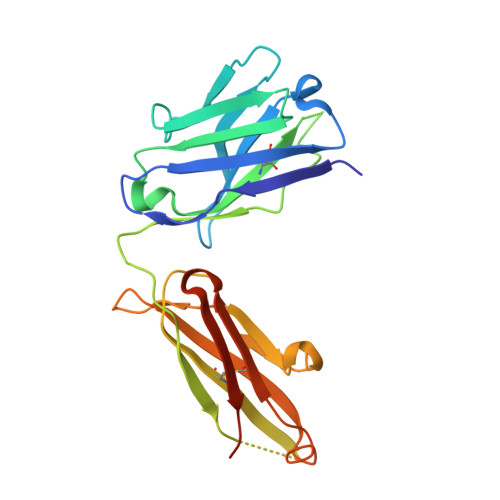
Human Antibody Bispecifics through Phage Display Selection.
Luthra, A., Langley, D.B., Schofield, P., Jackson, J., Abdelatti, M., Rouet, R., Nevoltris, D., Mazigi, O., Crossett, B., Christie, M., Christ, D.(2019) Biochemistry 58: 1701-1704
- PubMed: 30900875 Search on PubMed
- DOI: https://doi.org/10.1021/acs.biochem.9b00037
- Primary Citation Related Structures:
6MH2 - PubMed Abstract:
We developed a repertoire approach to generate human antibody bispecifics. Using phage display selection of antibody heavy chains in the presence of a competitor light chain and providing a cognate light chain with an affinity handle, we identified mutations that prevent heavy/light chain mispairing. The strategy allows for the selection of human antibody chains that autonomously assemble into bispecifics.
- Garvan Institute of Medical Research , 384 Victoria Road , Darlinghurst, Sydney , New South Wales 2010 , Australia.
Organizational Affiliation: