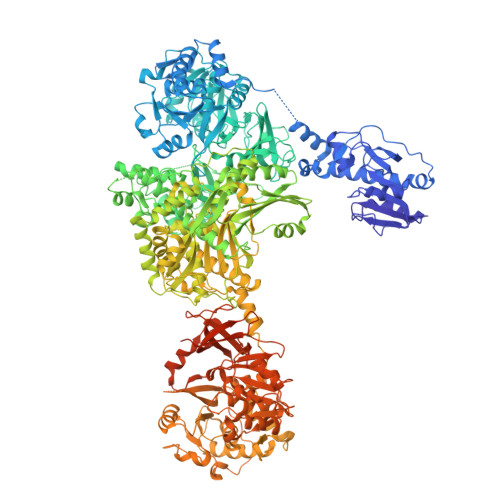
Structures of a dimodular nonribosomal peptide synthetase reveal conformational flexibility.
Reimer, J.M., Eivaskhani, M., Harb, I., Guarne, A., Weigt, M., Schmeing, T.M.(2019) Science 366
- PubMed: 31699907 Search on PubMed
- DOI: https://doi.org/10.1126/science.aaw4388
- Primary Citation Related Structures:
6MFW, 6MFX, 6MFY, 6MFZ, 6MG0 - PubMed Abstract:
Nonribosomal peptide synthetases (NRPSs) are biosynthetic enzymes that synthesize natural product therapeutics using a modular synthetic logic, whereby each module adds one aminoacyl substrate to the nascent peptide. We have determined five x-ray crystal structures of large constructs of the NRPS linear gramicidin synthetase, including a structure of a full core dimodule in conformations organized for the condensation reaction and intermodular peptidyl substrate delivery. The structures reveal differences in the relative positions of adjacent modules, which are not strictly coupled to the catalytic cycle and are consistent with small-angle x-ray scattering data. The structures and covariation analysis of homologs allowed us to create mutants that improve the yield of a peptide from a module-swapped dimodular NRPS.
- Department of Biochemistry and Center de Recherche en Biologie Structurale, McGill University, Montréal, QC H3G 0B1, Canada.
Organizational Affiliation: