Discriminative T-cell receptor recognition of highly homologous HLA-DQ2-bound gluten epitopes.
Dahal-Koirala, S., Ciacchi, L., Petersen, J., Risnes, L.F., Neumann, R.S., Christophersen, A., Lundin, K.E.A., Reid, H.H., Qiao, S.W., Rossjohn, J., Sollid, L.M.(2019) J Biological Chem 294: 941-952
- PubMed: 30455354 Search on PubMedSearch on PubMed Central
- DOI: https://doi.org/10.1074/jbc.RA118.005736
- Primary Citation Related Structures:
6MFF, 6MFG - PubMed Abstract:
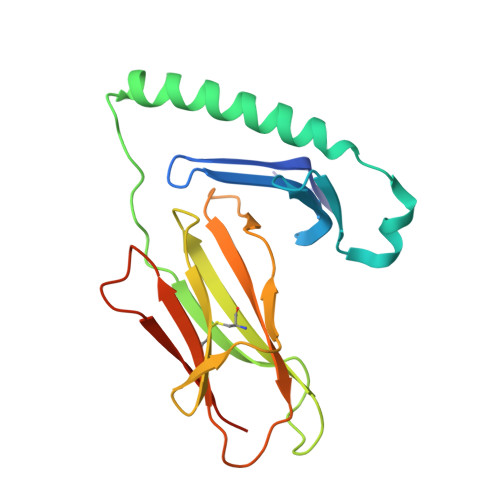
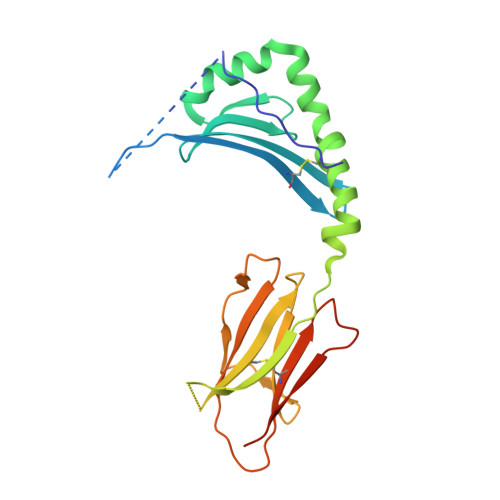
Celiac disease (CeD) provides an opportunity to study the specificity underlying human T-cell responses to an array of similar epitopes presented by the same human leukocyte antigen II (HLA-II) molecule. Here, we investigated T-cell responses to the two immunodominant and highly homologous HLA-DQ2.5-restricted gluten epitopes, DQ2.5-glia-α1a (PFPQPELPY) and DQ2.5-glia-ω1 (PFPQPEQPF). Using HLA-DQ2.5-DQ2.5-glia-α1a and HLA-DQ2.5-DQ2.5-glia-ω1 tetramers and single-cell αβ T-cell receptor (TCR) sequencing, we observed that despite similarity in biased variable-gene usage in the TCR repertoire responding to these nearly identical peptide-HLA-II complexes, most of the T cells are specific for either of the two epitopes. To understand the molecular basis of this exquisite fine specificity, we undertook Ala substitution assays revealing that the p7 residue (Leu/Gln) is critical for specific epitope recognition by both DQ2.5-glia-α1a- and DQ2.5-glia-ω1-reactive T-cell clones. We determined high-resolution binary crystal structures of HLA-DQ2.5 bound to DQ2.5-glia-α1a (2.0 Å) and DQ2.5-glia-ω1 (2.6 Å). These structures disclosed that differences around the p7 residue subtly alter the neighboring substructure and electrostatic properties of the HLA-DQ2.5-peptide complex, providing the fine specificity underlying the responses against these two highly homologous gluten epitopes. This study underscores the ability of TCRs to recognize subtle differences in the peptide-HLA-II landscape in a human disease setting.
- From the Department of Immunology, University of Oslo and Oslo University Hospital-Rikshospitalet, 0372 Oslo, Norway.
Organizational Affiliation: