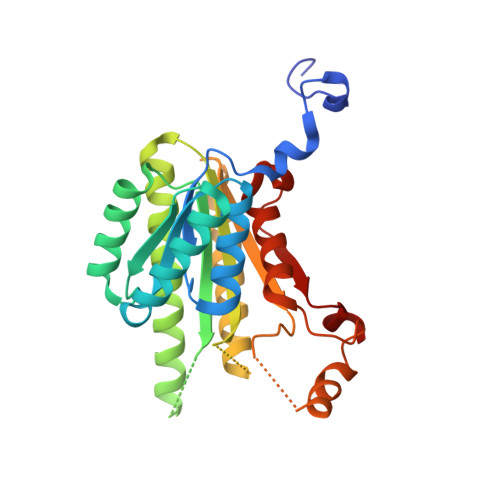
Structural and biochemical characterization of 20 beta-hydroxysteroid dehydrogenase fromBifidobacterium adolescentisstrain L2-32.
Doden, H.L., Pollet, R.M., Mythen, S.M., Wawrzak, Z., Devendran, S., Cann, I., Koropatkin, N.M., Ridlon, J.M.(2019) J Biological Chem 294: 12040-12053
- PubMed: 31209107 Search on PubMedSearch on PubMed Central
- DOI: https://doi.org/10.1074/jbc.RA119.009390
- Primary Citation Related Structures:
6M9U, 6OW4 - PubMed Abstract:
Anaerobic bacteria inhabiting the human gastrointestinal tract have evolved various enzymes that modify host-derived steroids. The bacterial steroid-17,20-desmolase pathway cleaves the cortisol side chain, forming pro-androgens predicted to impact host physiology. Bacterial 20β-hydroxysteroid dehydrogenase (20β-HSDH) regulates cortisol side-chain cleavage by reducing the C-20 carboxyl group on cortisol, yielding 20β-dihydrocortisol. Recently, the gene encoding 20β-HSDH in Butyricicoccus desmolans ATCC 43058 was reported, and a nonredundant protein search yielded a candidate 20 β- HSDH gene in Bifidobacterium adolescentis strain L2-32. B. adolescentis 20β-HSDH could regulate cortisol side-chain cleavage by limiting pro-androgen formation in bacteria such as Clostridium scindens and 21-dehydroxylation by Eggerthella lenta Here, the putative B. adolescentis 20β-HSDH was cloned, overexpressed, and purified. 20β-HSDH activity was confirmed through whole-cell and pure enzymatic assays, and it is specific for cortisol. Next, we solved the structures of recombinant 20β-HSDH in both the apo- and holo-forms at 2.0-2.2 Å resolutions, revealing close overlap except for rearrangements near the active site. Interestingly, the structures contain a large, flexible N-terminal region that was investigated by gel-filtration chromatography and CD spectroscopy. This extended N terminus is important for protein stability because deletions of varying lengths caused structural changes and reduced enzymatic activity. A nonconserved extended N terminus was also observed in several short-chain dehydrogenase/reductase family members. B. adolescentis strains capable of 20β-HSDH activity could alter glucocorticoid metabolism in the gut and thereby serve as potential probiotics for the management of androgen-dependent diseases.
- Microbiome Metabolic Engineering Theme, Carl R. Woese Institute for Genomic Biology, Urbana, Illinois 61801; Department of Animal Sciences, University of Illinois at Urbana-Champaign, Urbana, Illinois 61801.
Organizational Affiliation: