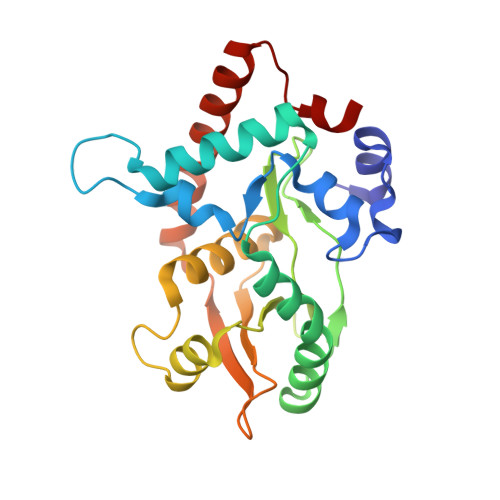
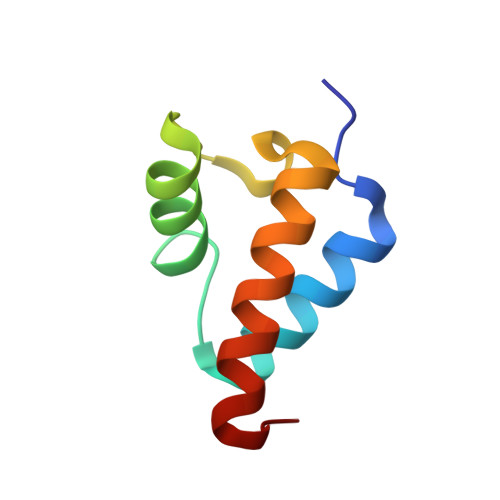
Crystal structure of the Red beta C-terminal domain in complex with lambda Exonuclease reveals an unexpected homology with lambda Orf and an interaction with Escherichia coli single stranded DNA binding protein.
Caldwell, B.J., Zakharova, E., Filsinger, G.T., Wannier, T.M., Hempfling, J.P., Chun-Der, L., Pei, D., Church, G.M., Bell, C.E.(2019) Nucleic Acids Res 47: 1950-1963
- PubMed: 30624736 Search on PubMedSearch on PubMed Central
- DOI: https://doi.org/10.1093/nar/gky1309
- Primary Citation Related Structures:
6M9K - PubMed Abstract:
Bacteriophage λ encodes a DNA recombination system that includes a 5'-3' exonuclease (λ Exo) and a single strand annealing protein (Redβ). The two proteins form a complex that is thought to mediate loading of Redβ directly onto the single-stranded 3'-overhang generated by λ Exo. Here, we present a 2.3 Å crystal structure of the λ Exo trimer bound to three copies of the Redβ C-terminal domain (CTD). Mutation of residues at the hydrophobic core of the interface disrupts complex formation in vitro and impairs recombination in vivo. The Redβ CTD forms a three-helix bundle with unexpected structural homology to phage λ Orf, a protein that binds to E. coli single-stranded DNA binding protein (SSB) to function as a recombination mediator. Based on this relationship, we found that Redβ binds to full-length SSB, and to a peptide corresponding to its nine C-terminal residues, in an interaction that requires the CTD. These results suggest a dual role of the CTD, first in binding to λ Exo to facilitate loading of Redβ directly onto the initial single-stranded DNA (ssDNA) at a 3'-overhang, and second in binding to SSB to facilitate annealing of the overhang to SSB-coated ssDNA at the replication fork.
- Ohio State Biochemistry Program, The Ohio State University, 484 West 12th Avenue, Columbus, OH 43210, USA.
Organizational Affiliation: