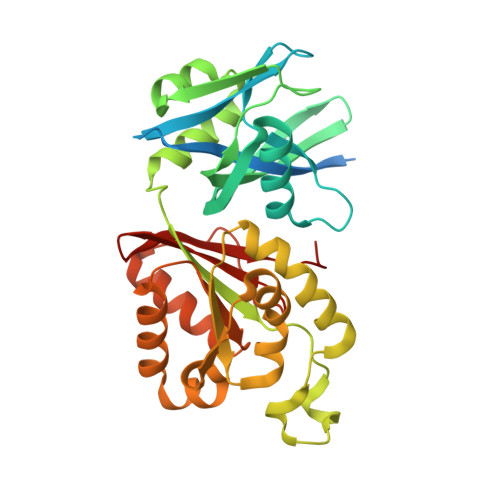
Plasticity, ligand conformation and enzyme action of Mycobacterium smegmatis MutT1.
Raj, P., Karthik, S., Arif, S.M., Varshney, U., Vijayan, M.(2020) Acta Crystallogr D Struct Biol 76: 982-992
- PubMed: 33021500 Search on PubMed
- DOI: https://doi.org/10.1107/S2059798320010992
- Primary Citation Related Structures:
6M65, 6M69, 6M6Y, 6M72 - PubMed Abstract:
Mycobacterium smegmatis MutT1 (MsMutT1) is a sanitation enzyme made up of an N-terminal Nudix hydrolase domain and a C-terminal domain resembling a histidine phosphatase. It has been established that the action of MutT1 on 8-oxo-dGTP, 8-oxo-GTP and diadenosine polyphosphates is modulated by intermolecular interactions. In order to further explore this and to elucidate the structural basis of its differential action on 8-oxo-NTPs and unsubstituted NTPs, the crystal structures of complexes of MsMutT1 with 8-oxo-dGTP, GMPPNP and GMPPCP have been determined. Replacement soaking was used in order to ensure that the complexes were isomorphous to one another. Analysis of the structural data led to the elucidation of a relationship between the arrangements of molecules observed in the crystals, molecular plasticity and the action of the enzyme on nucleotides. The dominant mode of arrangement involving a head-to-tail sequence predominantly leads to the generation of NDPs. The other mode of packing arrangement appears to preferentially generate NMPs. This work also provides interesting insights into the dependence of enzyme action on the conformation of the ligand. The possibility of modulating the enzyme action through differences in intermolecular interactions and ligand conformations makes MsMutT1 a versatile enzyme.
- Molecular Biophysics Unit, Indian Institute of Science, Bengaluru 560 012, India.
Organizational Affiliation: