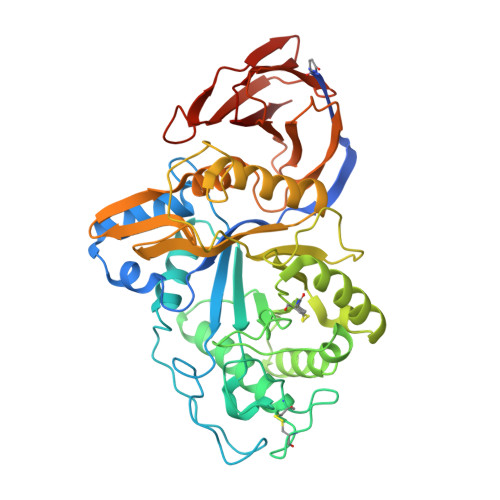
Crystal structure of GH30-7 endoxylanase C from the filamentous fungus Talaromyces cellulolyticus.
Nakamichi, Y., Fujii, T., Watanabe, M., Matsushika, A., Inoue, H.(2020) Acta Crystallogr F Struct Biol Commun 76: 341-349
- PubMed: 32744245 Search on PubMedSearch on PubMed Central
- DOI: https://doi.org/10.1107/S2053230X20009024
- Primary Citation Related Structures:
6M5Z - PubMed Abstract:
GH30-7 endoxylanase C from the cellulolytic fungus Talaromyces cellulolyticus (TcXyn30C) belongs to glycoside hydrolase family 30 subfamily 7, and specifically releases 2 2 -(4-O-methyl-α-D-glucuronosyl)-xylobiose from glucuronoxylan, as well as various arabino-xylooligosaccharides from arabinoxylan. TcXyn30C has a modular structure consisting of a catalytic domain and a C-terminal cellulose-binding module 1 (CBM1). In this study, the crystal structure of a TcXyn30C mutant which lacks the CBM1 domain was determined at 1.65 Å resolution. The structure of the active site of TcXyn30C was compared with that of the bifunctional GH30-7 xylanase B from T. cellulolyticus (TcXyn30B), which exhibits glucuronoxylanase and xylobiohydrolase activities. The results revealed that TcXyn30C has a conserved structural feature for recognizing the 4-O-methyl-α-D-glucuronic acid (MeGlcA) substituent in subsite -2b. Additionally, the results demonstrated that Phe47 contributes significantly to catalysis by TcXyn30C. Phe47 is located in subsite -2b and also near the C-3 hydroxyl group of a xylose residue in subsite -2a. Substitution of Phe47 with an arginine residue caused a remarkable decrease in the catalytic efficiency towards arabinoxylan, suggesting the importance of Phe47 in arabinoxylan hydrolysis. These findings indicate that subsite -2b of TcXyn30C has unique structural features that interact with arabinofuranose and MeGlcA substituents.
- Bioconversion Group, Research Institute for Sustainable Chemistry, National Institute of Advanced Industrial Science and Technology (AIST), 3-11-32 Kagamiyama, Higashi-Hiroshima, Hiroshima 739-0046, Japan.
Organizational Affiliation: