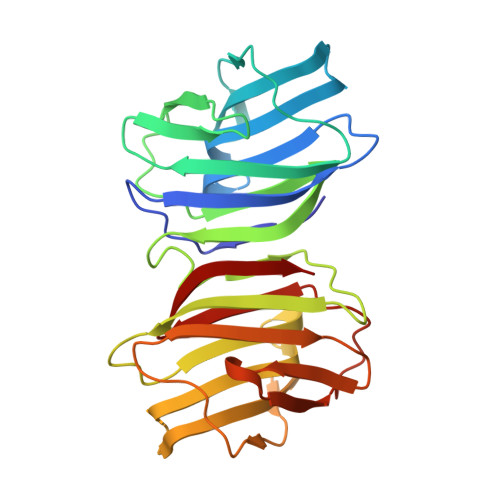
Crystal structure and conformational stability of a galectin-1 tandem-repeat mutant with a short linker.
Nonaka, Y., Ogawa, T., Shoji, H., Nishi, N., Kamitori, S., Nakamura, T.(2021) Glycobiology
- PubMed: 34735570 Search on PubMed
- DOI: https://doi.org/10.1093/glycob/cwab101
- Primary Citation Related Structures:
6M5Y - PubMed Abstract:
Modification of the domain architecture of galectins has been attempted to analyze their biological functions and to develop medical applications. Several types of galectin-1 repeat mutants were previously reported but, however, it was not clear whether the native structure of the wild type was retained. In this study, we determined the crystal structure of a galectin-1 tandem-repeat mutant with a short linker peptide, and compared the unfolding profiles of the wild type and mutant by chemical denaturation. The structure of the mutant was consistent with that of the dimer of the wild type, and both carbohydrate-binding sites were retained. The unfolding curve of the wild type with lactose suggested that the dimer dissociation and the tertiary structure unfolding was concomitant at micromolar protein concentrations. The midpoint denaturant concentration of the wild type was dependent on the protein concentration and lower than that of the mutant. Linking the two subunits significantly stabilized the tertiary structure. The mutant exhibited higher T-cell growth-inhibition activity and comparable hemagglutinating activity. Structural stabilization may prevent the oxidation of the internal cysteine residue.
- Department of Endocrinology, Faculty of Medicine, Kagawa University, 1750-1, Ikenobe, Miki-cho, Kita-gun, Kagawa 761-0793, Japan.
Organizational Affiliation: