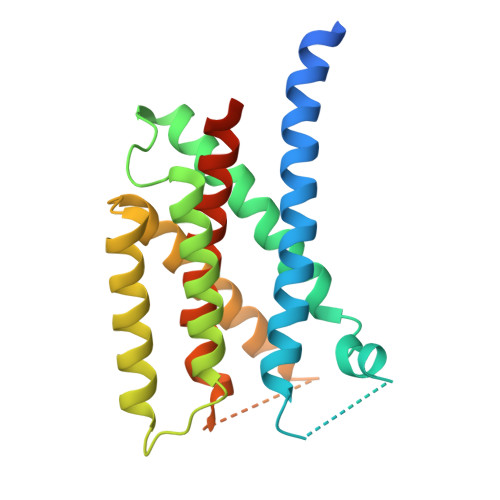
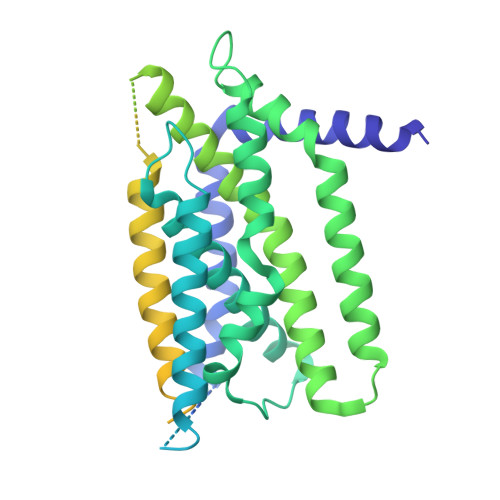
A structure of human Scap bound to Insig-2 suggests how their interaction is regulated by sterols.
Yan, R., Cao, P., Song, W., Qian, H., Du, X., Coates, H.W., Zhao, X., Li, Y., Gao, S., Gong, X., Liu, X., Sui, J., Lei, J., Yang, H., Brown, A.J., Zhou, Q., Yan, C., Yan, N.(2021) Science 371
- PubMed: 33446483 Search on PubMed
- DOI: https://doi.org/10.1126/science.abb2224
- Primary Citation Related Structures:
6M49 - PubMed Abstract:
The sterol regulatory element-binding protein (SREBP) pathway controls cellular homeostasis of sterols. The key players in this pathway, Scap and Insig-1 and -2, are membrane-embedded sterol sensors. The 25-hydroxycholesterol (25HC)-dependent association of Scap and Insig acts as the master switch for the SREBP pathway. Here, we present cryo-electron microscopy analysis of the human Scap and Insig-2 complex in the presence of 25HC, with the transmembrane (TM) domains determined at an average resolution of 3.7 angstrom. The sterol-sensing domain in Scap and all six TMs in Insig-2 were resolved. A 25HC molecule is sandwiched between the S4 to S6 segments in Scap and TMs 3 and 4 in Insig-2 in the luminal leaflet of the membrane. Unwinding of the middle of the Scap-S4 segment is crucial for 25HC binding and Insig association.
- Westlake Laboratory of Life Sciences and Biomedicine, Key Laboratory of Structural Biology of Zhejiang Province, School of Life Sciences, Westlake University, Hangzhou 310024, Zhejiang Province, China.
Organizational Affiliation: