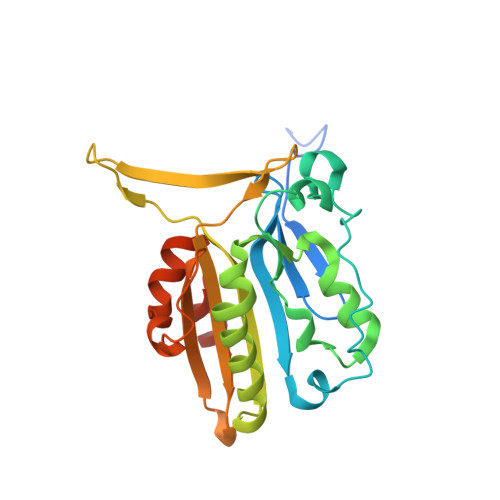
Crystal structure of the mouse endonuclease G.
Park, K.H., Yoon, S.M., Song, H.N., Yang, J.H., Ryu, S.E., Woo, E.J.(2020) Biochem Biophys Res Commun 526: 35-40
- PubMed: 32192768 Search on PubMed
- DOI: https://doi.org/10.1016/j.bbrc.2020.03.060
- Primary Citation Related Structures:
6LYF, 6M3F, 6M3T - PubMed Abstract:
Endonuclease G (EndoG) is a mitochondrial enzyme that responds to apoptotic stimuli by translocating to the nucleus and cleaving the chromatin DNA. The molecular mechanism of EndoG still remains unknown in higher organisms. Here, we determined the crystal structure of mouse EndoG at ∼1.96 Å resolution. The EndoG shows an altered dimeric configuration in which N-terminal region of one subunit interact to the other subunit in dimer. The deletion of this region that is highly conserved in mammalian EndoGs resulted in a monomer with significantly reduced activity suggesting the association of the dimeric arrangement into the nuclease activity. Furthermore, we observed a large conformational change in the loop of the active site groove in EndoG, which corresponds to the DNA binding region. Intriguingly, EndoG dimers are linked by oxidation of the reactive cysteine 110 in this flexible loop to form a long oligomeric chain in the crystal lattice. The structural analysis and ensuing biochemical data suggest that this flexible loop region in the active site is important to the regulation of EndoG nuclease function in mouse.
- Disease Target Structure Research Center, Korea Research Institute of Bioscience & Biotechnology (KRIBB), Daejeon, 305-806, Republic of Korea.
Organizational Affiliation: