Structural insights into the regulation of SigB activity by RsbV and RsbW.
Pathak, D., Jin, K.S., Tandukar, S., Kim, J.H., Kwon, E., Kim, D.Y.(2020) IUCrJ 7: 737-747
Experimental Data Snapshot
Starting Models: experimental
View more details
wwPDB Validation 3D Report Full Report
(2020) IUCrJ 7: 737-747
Macromolecule Content
Entity ID: 1 | |||||
|---|---|---|---|---|---|
| Molecule | Chains | Sequence Length | Organism | Details | Image |
| Serine-protein kinase RsbW | 141 | Bacillus subtilis subsp. subtilis str. 168 | Mutation(s): 0 Gene Names: rsbW, BSU04720 EC: 2.7.11.1 | 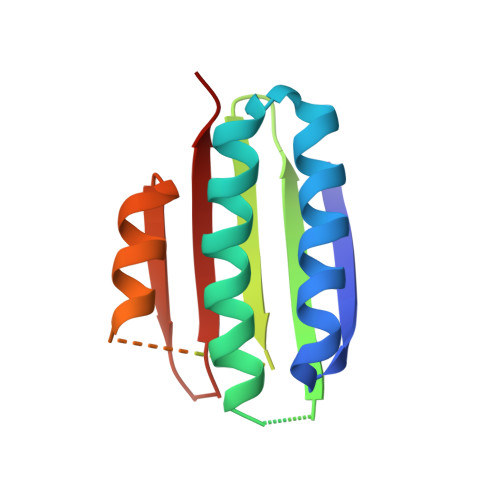 | |
UniProt | |||||
Entity Groups | |||||
| Sequence Clusters | 30% Identity50% Identity70% Identity90% Identity95% Identity100% Identity | ||||
| UniProt Group | P17904 | ||||
Sequence AnnotationsExpand | |||||
Reference Sequence | |||||
Entity ID: 2 | |||||
|---|---|---|---|---|---|
| Molecule | Chains | Sequence Length | Organism | Details | Image |
| Anti-sigma-B factor antagonist | 103 | Bacillus subtilis subsp. subtilis str. 168 | Mutation(s): 0 Gene Names: rsbV, BSU04710 | 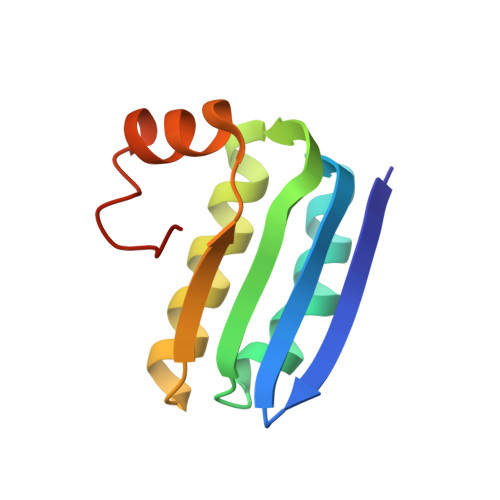 | |
UniProt | |||||
Entity Groups | |||||
| Sequence Clusters | 30% Identity50% Identity70% Identity90% Identity95% Identity100% Identity | ||||
| UniProt Group | P17903 | ||||
Sequence AnnotationsExpand | |||||
Reference Sequence | |||||
| Length ( Å ) | Angle ( ˚ ) |
|---|---|
| a = 117.23 | α = 90 |
| b = 70.94 | β = 105.35 |
| c = 137.68 | γ = 90 |
| Software Name | Purpose |
|---|---|
| PHENIX | refinement |
| PDB_EXTRACT | data extraction |
| MOSFLM | data reduction |
| Aimless | data scaling |
| PHASER | phasing |
| Funding Organization | Location | Grant Number |
|---|---|---|
| National Research Foundation (NRF, Korea) | Korea, Republic Of | 2017R1D1A1B03034088 |
| National Research Foundation (NRF, Korea) | Korea, Republic Of | 2017R1A6A3A11029218 |