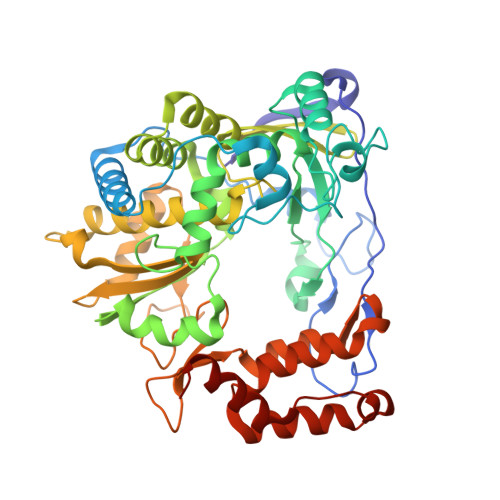
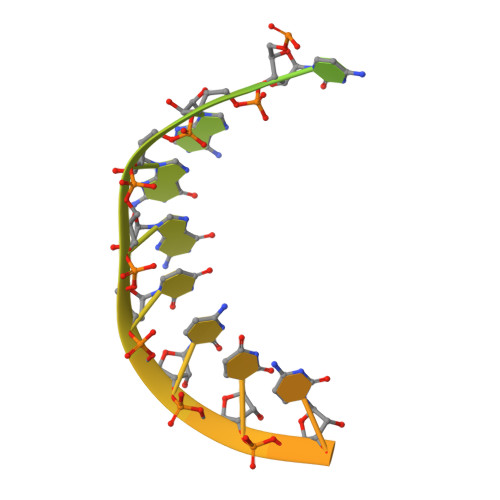
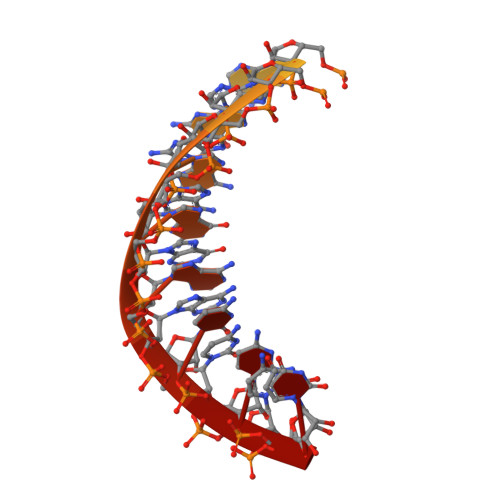
Stringent control of the RNA-dependent RNA polymerase translocation revealed by multiple intermediate structures.
Wang, M., Li, R., Shu, B., Jing, X., Ye, H.Q., Gong, P.(2020) Nat Commun 11: 2605-2605
- PubMed: 32451382 Search on PubMedSearch on PubMed Central
- DOI: https://doi.org/10.1038/s41467-020-16234-4
- Primary Citation Related Structures:
6LSE, 6LSF, 6LSG, 6LSH - PubMed Abstract:
Each polymerase nucleotide addition cycle is associated with two primary conformational changes of the catalytic complex: the pre-chemistry active site closure and post-chemistry translocation. While active site closure is well interpreted by numerous crystallographic snapshots, translocation intermediates are rarely captured. Here we report three types of intermediate structures in an RNA-dependent RNA polymerase (RdRP). The first two types, captured in forward and reverse translocation events, both highlight the role of RdRP-unique motif G in restricting the RNA template movement, corresponding to the rate-limiting step in translocation. By mutating two critical residues in motif G, we obtain the third type of intermediates that may mimic the transition state of this rate-limiting step, demonstrating a previously unidentified movement of the template strand. We propose that a similar strategy may be utilized by other classes of nucleic acid polymerases to ensure templating nucleotide positioning for efficient catalysis through restricting interactions with template RNA.
- Key Laboratory of Special Pathogens and Biosafety, Wuhan Institute of Virology, Center for Biosafety Mega-Science, Chinese Academy of Sciences, No.44 Xiao Hong Shan, Wuhan, Hubei, 430071, China.
Organizational Affiliation: