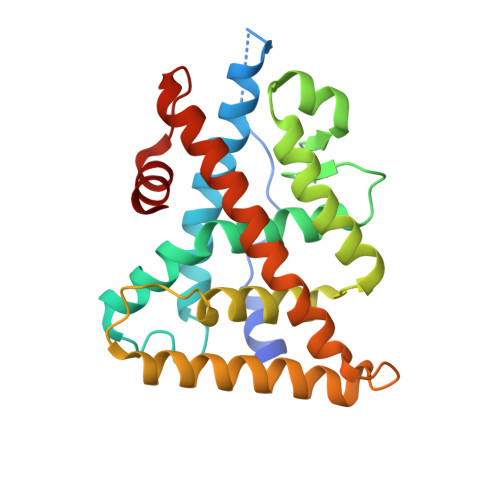
Structural Insights into the Specificity of Ligand Binding and Coactivator Assembly by Estrogen-Related Receptor beta.
Yao, B., Zhang, S., Wei, Y., Tian, S., Lu, Z., Jin, L., He, Y., Xie, W., Li, Y.(2020) J Mol Biology 432: 5460-5472
- PubMed: 32795533 Search on PubMed
- DOI: https://doi.org/10.1016/j.jmb.2020.08.007
- Primary Citation Related Structures:
6LIT, 6LN4 - PubMed Abstract:
Estrogen-related receptor β (ERRβ) is a nuclear receptor critical for many biological processes. Despite the biological and pharmaceutical importance of ERRβ, deciphering the structure of ERRβ has been hampered by the difficulties in obtaining a pure and stable protein for structural studies. In fact, the ERRβ ligand-binding domain remains the last unsolved ERR structure and also one of only a few unknown nuclear receptor structures. Here, we report the identification of a critical single-residue mutation resulted in robust solubility and stability of an active ERRβ ligand-binding domain, thereby providing a protein tool enabling the first probe into the biochemical and structural studies of this important receptor. The crystal structure reveals key structural features that have enabled the integration of the molecular determinants of signals transduced across the ligand binding and coregulator recruitment by all three ERR subtypes, which also provides a framework for the rational design of selective and potent ligands for the treatment of various ERR-mediated diseases.
- The State Key Laboratory of Cellular Stress Biology, Innovation Center for Cell Signaling Network, School of Life Sciences, Xiamen University, Fujian 361005, China.
Organizational Affiliation: