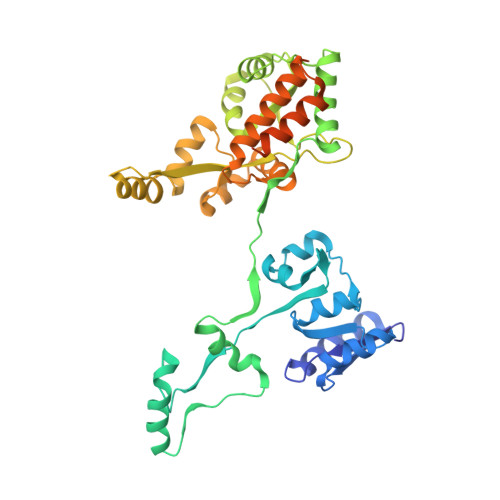
Passenger sequences can promote interlaced dimers in a common variant of the maltose-binding protein.
Momin, A.A., Hameed, U.F.S., Arold, S.T.(2019) Sci Rep 9: 20396-20396
- PubMed: 31892719 Search on PubMedSearch on PubMed Central
- DOI: https://doi.org/10.1038/s41598-019-56718-y
- Primary Citation Related Structures:
6LES, 6LF3 - PubMed Abstract:
The maltose-binding protein (MBP) is one of the most frequently used protein tags due to its capacity to stabilize, solubilize and even crystallize recombinant proteins that are fused to it. Given that MBP is thought to be a highly stable monomeric protein with known characteristics, fused passenger proteins are often studied without being cleaved from MBP. Here we report that a commonly used engineered MBP version (mutated to lower its surface entropy) can form interlaced dimers when fused to short protein sequences derived from the focal adhesion kinase (FAK) or the homologous protein tyrosine kinase 2 (PYK2). These MBP dimers still bind maltose and can interconvert with monomeric forms in vitro under standard conditions despite a contact surface of more than 11,000 Å 2 . We demonstrate that both the mutations in MBP and the fused protein sequences were required for dimer formation. The FAK and PYK2 sequences are less than 40% identical, monomeric, and did not show specific interactions with MBP, suggesting that a variety of sequences can promote this MBP dimerization. MBP dimerization was abrogated by reverting two of the eight mutations introduced in the engineered MBP. Our results provide an extreme example for induced reversible domain-swapping, with implications for protein folding dynamics. Our observations caution that passenger-promoted MBP dimerization might mislead experimental characterization of the fused protein sequences, but also suggest a simple mutation to stop this phenomenon.
- King Abdullah University of Science and Technology (KAUST), Computational Bioscience Research Center (CBRC), Division of Biological and Environmental Sciences and Engineering (BESE), Thuwal, 23955-6900, Saudi Arabia.
Organizational Affiliation: