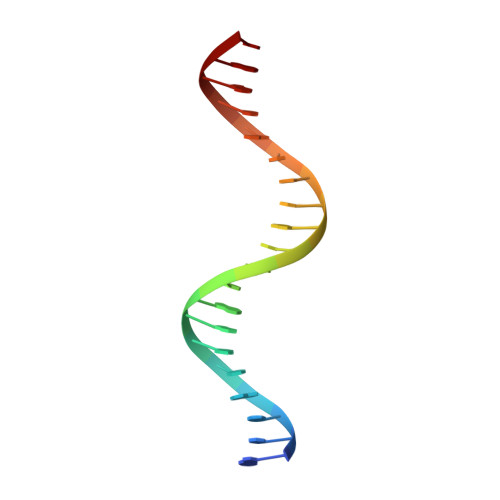
Mechanism of forkhead transcription factors binding to a novel palindromic DNA site.
Li, J., Dai, S., Chen, X., Liang, X., Qu, L., Jiang, L., Guo, M., Zhou, Z., Wei, H., Zhang, H., Chen, Z., Chen, L., Chen, Y.(2021) Nucleic Acids Res 49: 3573-3583
- PubMed: 33577686 Search on PubMedSearch on PubMed Central
- DOI: https://doi.org/10.1093/nar/gkab086
- Primary Citation Related Structures:
6LBI, 6LBM - PubMed Abstract:
Forkhead transcription factors bind a canonical consensus DNA motif, RYAAAYA (R = A/G, Y = C/T), as a monomer. However, the molecular mechanisms by which forkhead transcription factors bind DNA as a dimer are not well understood. In this study, we show that FOXO1 recognizes a palindromic DNA element DIV2, and mediates transcriptional regulation. The crystal structure of FOXO1/DIV2 reveals that the FOXO1 DNA binding domain (DBD) binds the DIV2 site as a homodimer. The wing1 region of FOXO1 mediates the dimerization, which enhances FOXO1 DNA binding affinity and complex stability. Further biochemical assays show that FOXO3, FOXM1 and FOXI1 also bind the DIV2 site as homodimer, while FOXC2 can only bind this site as a monomer. Our structural, biochemical and bioinformatics analyses not only provide a novel mechanism by which FOXO1 binds DNA as a homodimer, but also shed light on the target selection of forkhead transcription factors.
- Department of Oncology, NHC Key Laboratory of Cancer Proteomics, Laboratory of Structural Biology, National Clinical Research Center for Geriatric Disorders, Xiangya Hospital, Central South University, Changsha, Hunan 410008, China.
Organizational Affiliation: