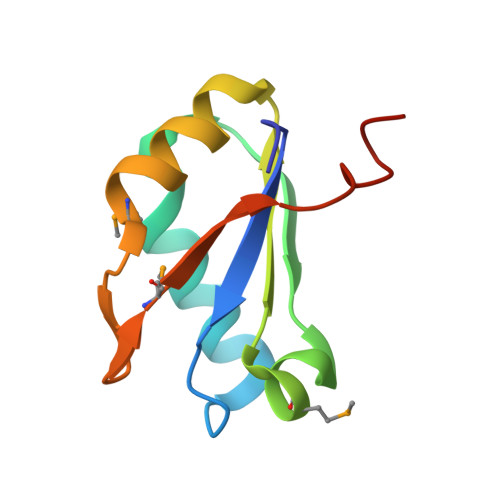
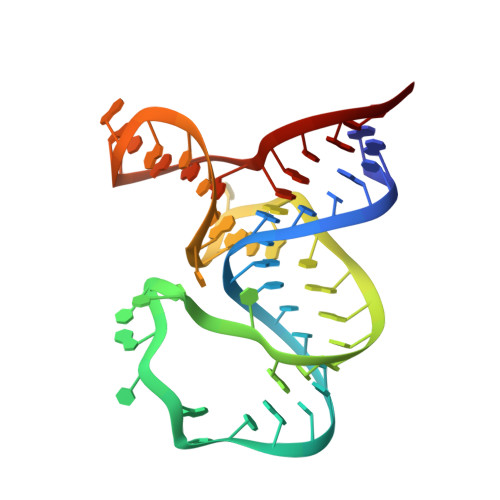
SAM-VI riboswitch structure and signature for ligand discrimination.
Sun, A., Gasser, C., Li, F., Chen, H., Mair, S., Krasheninina, O., Micura, R., Ren, A.(2019) Nat Commun 10: 5728-5728
- PubMed: 31844059 Search on PubMedSearch on PubMed Central
- DOI: https://doi.org/10.1038/s41467-019-13600-9
- Primary Citation Related Structures:
6LAS, 6LAU, 6LAX, 6LAZ - PubMed Abstract:
Riboswitches are metabolite-sensing, conserved domains located in non-coding regions of mRNA that are central to regulation of gene expression. Here we report the first three-dimensional structure of the recently discovered S-adenosyl-L-methionine responsive SAM-VI riboswitch. SAM-VI adopts a unique fold and ligand pocket that are distinct from all other known SAM riboswitch classes. The ligand binds to the junctional region with its adenine tightly intercalated and Hoogsteen base-paired. Furthermore, we reveal the ligand discrimination mode of SAM-VI by additional X-ray structures of this riboswitch bound to S-adenosyl-L-homocysteine and a synthetic ligand mimic, in combination with isothermal titration calorimetry and fluorescence spectroscopy to explore binding thermodynamics and kinetics. The structure is further evaluated by analysis of ligand binding to SAM-VI mutants. It thus provides a thorough basis for developing synthetic SAM cofactors for applications in chemical and synthetic RNA biology.
- Life Sciences Institute, Zhejiang University, 310058, Hangzhou, Zhejiang, China.
Organizational Affiliation: