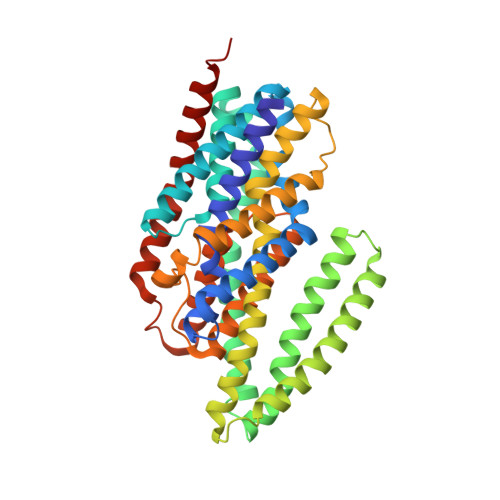
Structure of the sodium-dependent phosphate transporter reveals insights into human solute carrier SLC20.
Tsai, J.-Y., Chu, C.-H., Lin, M.-G., Chou, Y.-H., Hong, R.-Y., Yen, C.-Y., Hsiao, C.-D., Sun, Y.-J.(2020) Sci Adv 6: eabb4024-eabb4024
- PubMed: 32821837 Search on PubMedSearch on PubMed Central
- DOI: https://doi.org/10.1126/sciadv.abb4024
- Primary Citation Related Structures:
6L85 - PubMed Abstract:
Inorganic phosphate (P i ) is a fundamental and essential element for nucleotide biosynthesis, energy supply, and cellular signaling in living organisms. Human phosphate transporter ( h PiT) dysfunction causes numerous diseases, but the molecular mechanism underlying transporters remains elusive. We report the structure of the sodium-dependent phosphate transporter from Thermotoga maritima ( Tm PiT) in complex with sodium and phosphate ( Tm PiT-Na/Pi) at 2.3-angstrom resolution. We reveal that one phosphate and two sodium ions (Pi-2Na) are located at the core of Tm PiT and that the third sodium ion (Na fore ) is located near the inner membrane boundary. We propose an elevator-like mechanism for sodium and phosphate transport by Tm PiT, with the Tm PiT-Na/Pi complex adopting an inward occluded conformation. We found that disease-related h PiT variants carry mutations in the corresponding sodium- and phosphate-binding residues identified in Tm PiT. Our three-dimensional structure of Tm PiT provides a framework for understanding PiT dysfunction and for future structure-based drug design.
- Department of Life Science and Institute of Bioinformatics and Structural Biology, College of Life Science, National Tsing Hua University, Hsinchu 30013, Taiwan (R.O.C.).
Organizational Affiliation: