A structural study of TatD from Staphylococcus aureus elucidates a putative DNA-binding mode of a Mg2+-dependent nuclease.
Lee, K.-Y., Cheon, S.-H., Kim, D.-G., Lee, S.J., Lee, B.-J.(2020) IUCrJ 7: 509-521
Experimental Data Snapshot
(2020) IUCrJ 7: 509-521
Macromolecule Content
Entity ID: 1 | |||||
|---|---|---|---|---|---|
| Molecule | Chains | Sequence Length | Organism | Details | Image |
| Deoxyribonuclease YcfH | 273 | Staphylococcus aureus | Mutation(s): 0 Gene Names: yabD, tatP, ycfH, ycfH_1, BN1321_120011, C7P97_09010, CSC83_12190, CSC87_02770, EP54_03105, EQ90_10980... EC: 3.1.21 | 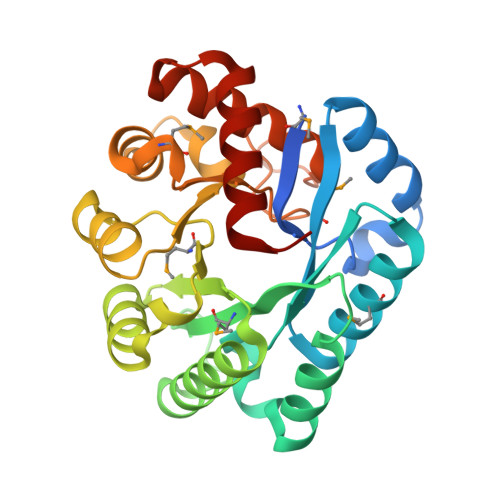 | |
UniProt | |||||
Entity Groups | |||||
| Sequence Clusters | 30% Identity50% Identity70% Identity90% Identity95% Identity100% Identity | ||||
| UniProt Group | W8U6D8 | ||||
Sequence AnnotationsExpand | |||||
Reference Sequence | |||||
| Ligands 2 Unique | |||||
|---|---|---|---|---|---|
| ID | Chains | Name / Formula / InChI Key | 2D Diagram | 3D Interactions | |
| PO4 (Subject of Investigation/LOI) Download:Ideal Coordinates CCD File | E [auth A], F [auth A], I [auth B], J [auth B] | PHOSPHATE ION O4 P NBIIXXVUZAFLBC-UHFFFAOYSA-K |  | ||
| NI (Subject of Investigation/LOI) Download:Ideal Coordinates CCD File | C [auth A], D [auth A], G [auth B], H [auth B] | NICKEL (II) ION Ni VEQPNABPJHWNSG-UHFFFAOYSA-N |  | ||
| Modified Residues 1 Unique | |||||
|---|---|---|---|---|---|
| ID | Chains | Type | Formula | 2D Diagram | Parent |
| MSE Query on MSE | A, B | L-PEPTIDE LINKING | C5 H11 N O2 Se |  | MET |
| Length ( Å ) | Angle ( ˚ ) |
|---|---|
| a = 47.627 | α = 90 |
| b = 77.832 | β = 98.81 |
| c = 77.378 | γ = 90 |
| Software Name | Purpose |
|---|---|
| REFMAC | refinement |
| PDB_EXTRACT | data extraction |
| HKL-2000 | data reduction |
| HKL-2000 | data scaling |
| PHENIX | phasing |
| Funding Organization | Location | Grant Number |
|---|---|---|
| National Research Foundation (NRF, Korea) | Korea, Republic Of | NRF-2018R1A2A1A19018526 |
| National Research Foundation (NRF, Korea) | Korea, Republic Of | NRF-2018R1A5A2024425 |
| National Research Foundation (NRF, Korea) | Korea, Republic Of | NRF-2016R1C1B2014609 |
| National Research Foundation (NRF, Korea) | Korea, Republic Of | NRF-2019R1H1A1102102 |