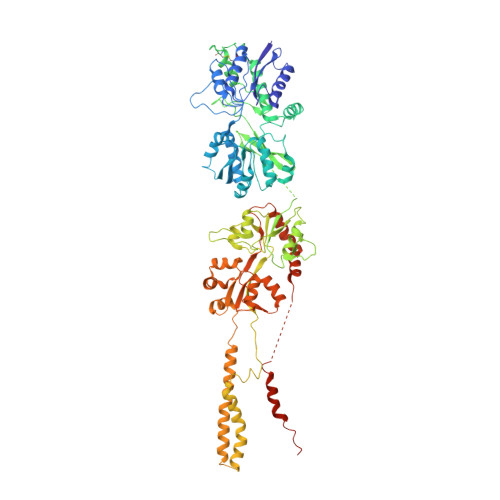
Structural dynamics of the GluK3-kainate receptor neurotransmitter binding domains revealed by cryo-EM.
Kumari, J., Bendre, A.D., Bhosale, S., Vinnakota, R., Burada, A.P., Tria, G., Ravelli, R.B.G., Peters, P.J., Joshi, M., Kumar, J.(2020) Int J Biol Macromol 149: 1051-1058
- PubMed: 32006583 Search on PubMed
- DOI: https://doi.org/10.1016/j.ijbiomac.2020.01.282
- Primary Citation Related Structures:
6KZM, 6L6F - PubMed Abstract:
Kainate receptors belong to the ionotropic glutamate receptor family and play critical roles in the regulation of synaptic networks. The kainate receptor subunit GluK3 has unique functional properties and contributes to presynaptic facilitation at the hippocampal mossy fiber synapses along with roles at the post-synapses. To gain structural insights into the unique functional properties and dynamics of GluK3 receptor, we imaged them via electron microscopy in the apo-state and in complex with either agonist kainate or antagonist UBP301. Our analysis of all the GluK3 full-length structures not only provides insights into the receptor transitions between desensitized and closed states but also reveals a "non-classical" conformation of neurotransmitter binding domain in the closed-state distinct from that observed in AMPA and other kainate receptor structures. We show by molecular dynamics simulations that Asp759 influences the stability of the LBD dimers and hence could be responsible for the observed conformational variability and dynamics of the GluK3 via electron microscopy. Lower dimer stability could explain faster desensitization and low agonist sensitivity of GluK3. In overview, our work helps to associate biochemistry and physiology of GluK3 receptors with their structural biology and offers structural insights into the unique functional properties of these atypical receptors.
- Laboratory of Membrane Protein Biology, National Centre for Cell Science, NCCS Complex, S. P. Pune University, Pune, Maharashtra 411007, India.
Organizational Affiliation: