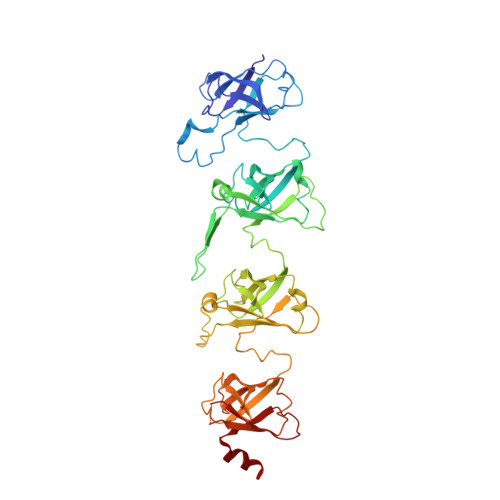
Cryo-EM Structure of a Bacterial Lipid Transporter YebT.
Liu, C., Ma, J., Wang, J., Wang, H.W., Zhang, L.(2020) J Mol Biology 432: 1008-1019
- PubMed: 31870848
- DOI: https://doi.org/10.1016/j.jmb.2019.12.008
- Primary Citation Related Structures:
6KZ3, 6KZ4 - PubMed Abstract:
The outer membrane (OM) of Gram-negative bacteria is asymmetric, with lipopolysaccharides (LPSs) on the outer surface and phospholipids (PLs) on the inner surface. This unique organization of OM makes Gram-negative bacteria resistant to many toxic chemicals. How this asymmetric distribution of lipids is maintained has been studied for decades with previous reports of an Mla (Maintenance of OM Lipid Asymmetry) system to be involved. Furthermore, the OM of Gram-negative bacteria is about 20 nm away from inner membrane (IM) where the lipids are synthesized. Therefore, how nascent lipids travel across the periplasmic space and arrive at the inner surface of OM is another interesting question. YebT is a homologue of MlaD in the Mla pathway, but its role in lipid distribution of the OM and IM is largely unknown. Here we report the first high-resolution (~3.0 Å) cryo-EM structure of full-length E. coli YebT in a substrate-bound state. Our structure with details of lipid interaction indicates that YebT is a lipid transporter spanning between IM and OM. We also demonstrate the symmetry mismatch in YebT and the existence of many other conformations of YebT revealing the intrinsic dynamics of this lipid channel. And a brief discussion on possible mechanisms of lipid transport is also included.
- Ministry of Education Key Laboratory of Protein Sciences, Tsinghua University, Beijing, China; Beijing Advanced Innovation Center for Structural Biology, Tsinghua University, Beijing, China; School of Life Sciences, Tsinghua University, Beijing, China; Tsinghua-Peking Joint Center for Life Sciences, Beijing, China.
Organizational Affiliation: