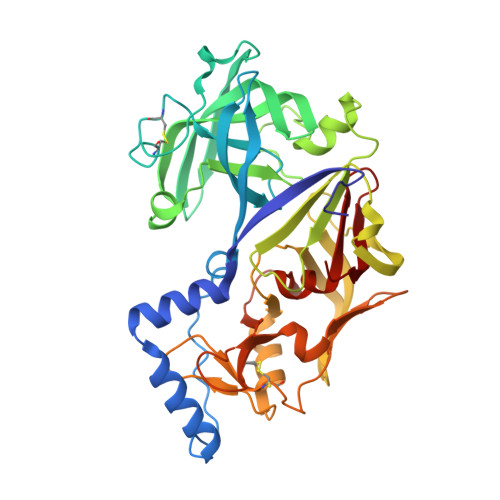
Activation mechanism of plasmepsins, pepsin-like aspartic proteases from Plasmodium, follows a unique trans-activation pathway.
Rathore, I., Mishra, V., Patel, C., Xiao, H., Gustchina, A., Wlodawer, A., Yada, R.Y., Bhaumik, P.(2021) FEBS J 288: 678-698
- PubMed: 32385863 Search on PubMedSearch on PubMed Central
- DOI: https://doi.org/10.1111/febs.15363
- Primary Citation Related Structures:
6KUB, 6KUC, 6KUD - PubMed Abstract:
Plasmodium parasites that cause malaria produce plasmepsins (PMs), pepsin-like aspartic proteases that are important antimalarial drug targets due to their role in host hemoglobin degradation. The enzymes are synthesized as inactive zymogens (pro-PMs), and the mechanism of their conversion to the active, mature forms has not been clearly elucidated. Our structural investigations of vacuolar pro-PMs with truncated prosegment (pro-tPMs) reveal that the formation of the S-shaped dimer is their innate property. Further structural studies, biochemical analysis, and molecular dynamics simulations indicate that disruption of the Tyr-Asp loop (121p-4), coordinated with the movement of the loop L1 (237-247) and helix H2 (101p-113p), is responsible for the extension of the pro-mature region (harboring the cleavage site). Consequently, under acidic pH conditions, these structural changes result in the dissociation of the dimers to monomers and the protonation of the residues in the prosegment prompts its unfolding. Subsequently, we demonstrated that the active site of the monomeric pro-tPMs with the unfolded prosegment is accessible for peptide substrate binding; in contrast, the active site is blocked in folded prosegment form of pro-tPMs. Thus, we propose a novel mechanism of auto-activation of vacuolar pro-tPMs that under acidic conditions can form a catalytically competent active site. One monomer cleaves the prosegment of the other one through a trans-activation process, resulting in formation of mature enzyme. As a result, once a mature enzyme is generated, it leads to the complete conversion of all the inactive pro-tPMs to their mature form. DATABASE: Atomic coordinates and structure factors have been submitted in the Protein Data Bank (PDB) under the PDB IDs 6KUB, 6KUC, and 6KUD.
- Department of Biosciences and Bioengineering, Indian Institute of Technology Bombay, Mumbai, India.
Organizational Affiliation: