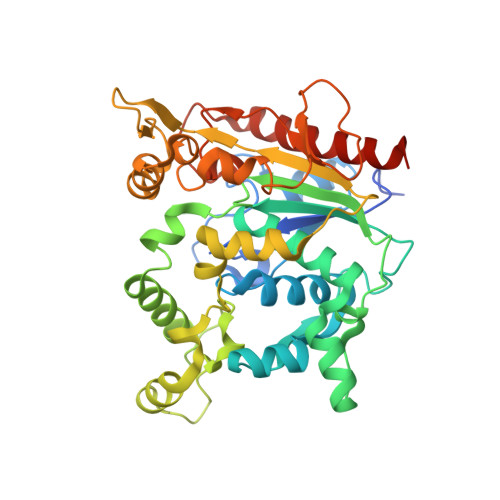
Crystal structure of pathogenic Staphylococcus aureus lipase complex with the anti-obesity drug orlistat.
Kitadokoro, K., Tanaka, M., Hikima, T., Okuno, Y., Yamamoto, M., Kamitani, S.(2020) Sci Rep 10: 5469-5469
- PubMed: 32214208 Search on PubMedSearch on PubMed Central
- DOI: https://doi.org/10.1038/s41598-020-62427-8
- Primary Citation Related Structures:
6KSI, 6KSL, 6KSM - PubMed Abstract:
Staphylococcus aureus lipase (SAL), a triacylglycerol esterase, is an important virulence factor and may be a therapeutic target for infectious diseases. Herein, we determined the 3D structure of native SAL, the mutated S116A inactive form, and the inhibitor complex using the anti-obesity drug orlistat to aid in drug development. The determined crystal structures showed a typical α/β hydrolase motif with a dimeric form. Fatty acids bound near the active site in native SAL and inactive S116A mutant structures. We found that orlistat potently inhibits SAL activity, and it covalently bound to the catalytic Ser116 residue. This is the first report detailing orlistat-lipase binding. It provides structure-based information on the production of potent anti-SAL drugs and lipase inhibitors. These results also indicated that orlistat can be repositioned to treat bacterial diseases.
- Faculty of Molecular Chemistry and Engineering, Graduate School of Science and Technology, Kyoto Institute of Technology, Hashigami-cho, Matsugasaki, Sakyo-ku, Kyoto, 606-8585, Japan. kengo@kit.ac.jp.
Organizational Affiliation: