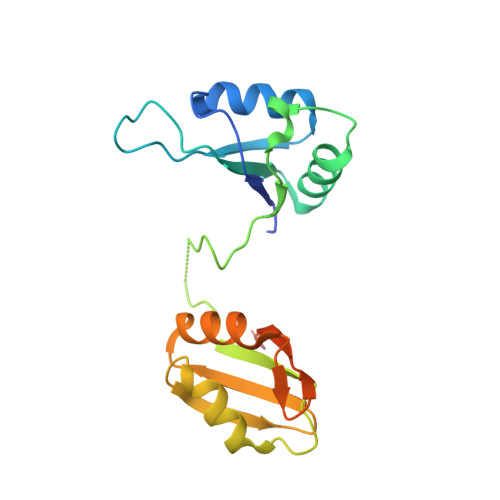
SYNCRIP, a new player in pri-let-7a processing.
Chen, Y., Chan, J., Chen, W., Li, J., Sun, M., Kannan, G.S., Mok, Y.K., Yuan, Y.A., Jobichen, C.(2020) RNA 26: 290-305
- PubMed: 31907208 Search on PubMedSearch on PubMed Central
- DOI: https://doi.org/10.1261/rna.072959.119
- Primary Citation Related Structures:
6KOR - PubMed Abstract:
microRNAs (miRNAs), a class of small and endogenous molecules that control gene expression, are broadly involved in biological processes. Although a number of cofactors that assist or antagonize let-7 miRNA biogenesis are well-established, more auxiliary factors remain to be investigated. Here, we identified SYNCRIP (Synaptotagmin Binding Cytoplasmic RNA Interacting Protein) as a new player for let-7a miRNA. SYNCRIP interacts with pri-let-7a both in vivo and in vitro. Knockdown of SYNCRIP impairs, while overexpression of SYNCRIP promotes, the expression of let-7a miRNA. A broad miRNA profiling analysis revealed that silencing of SYNCRIP regulates the expression of a set of mature miRNAs positively or negatively. In addition, SYNCRIP is associated with microprocessor complex and promotes the processing of pri-let-7a. Strikingly, the terminal loop of pri-let-7a was shown to be the main contributor for its interaction with SYNCRIP. Functional studies demonstrated that the SYNCRIP RRM2-3 domain can promote the processing of pri-let-7a. Structure-based alignment of RRM2-3 with other RNA binding proteins identified the residues likely to participate in protein-RNA interactions. Taken together, these findings suggest the promising role that SYNCRIP plays in miRNA regulation, thus providing insights into the function of SYNCRIP in eukaryotic development.
- Department of Biological Sciences and Centre for Bioimaging Sciences, National University of Singapore, Singapore 117543, Singapore.
Organizational Affiliation: