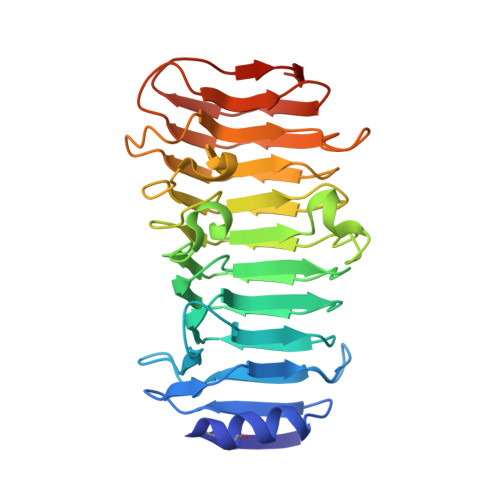
Structural and biochemical characterisation of a novel alginate lyase from Paenibacillus sp. str. FPU-7.
Itoh, T., Nakagawa, E., Yoda, M., Nakaichi, A., Hibi, T., Kimoto, H.(2019) Sci Rep 9: 14870-14870
- PubMed: 31619701 Search on PubMedSearch on PubMed Central
- DOI: https://doi.org/10.1038/s41598-019-51006-1
- Primary Citation Related Structures:
6KFN - PubMed Abstract:
A novel alginate lyase, PsAly, with a molecular mass of 33 kDa and whose amino acid sequence shares no significant similarity to other known proteins, was biochemically and structurally characterised from Paenibacillus sp. str. FPU-7. The maximum PsAly activity was obtained at 65 °C, with an optimum pH of pH 7-7.5. The activity was enhanced by divalent cations, such as Mg 2+ , Mn 2+ , or Co 2+ , and inhibited by a metal chelator, ethylenediaminetetraacetic acid. The reaction products indicated that PsAly is an endolytic enzyme with a preference for polymannuronate. Herein, we report a detailed crystal structure of PsAly at a resolution of 0.89 Å, which possesses a β-helix fold that creates a long cleft. The catalytic site was different from that of other polysaccharide lyases. Site-directed mutational analysis of conserved residues predicted Tyr184 and Lys221 as catalytic residues, abstracting from the C5 proton and providing a proton to the glycoside bond, respectively. One cation was found to bind to the bottom of the cleft and neutralise the carboxy group of the substrate, decreasing the pK a of the C5 proton to promote catalysis. Our study provides an insight into the structural basis for the catalysis of alginate lyases and β-helix polysaccharide lyases.
- Department of Bioscience and Biotechnology, Fukui Prefectural University, 4-1-1 Matsuoka Kenjyoujima, Eiheiji-cho, Yoshida-gun, Fukui, 910-1142, Japan. ito-t@fpu.ac.jp.
Organizational Affiliation: