The crystal structure of light-driven cyanobacterial chloride importer from Mastigocladopsis repens with Bromide ion
Yun, J.H., Park, J.H., Jin, Z., Ohki, M., Wang, Y., Lupala, C.S., Kim, M., Liu, H., Park, S.Y., Lee, W.To be published.
Experimental Data Snapshot
Starting Model: experimental
View more details
Macromolecule Content
Entity ID: 1 | |||||
|---|---|---|---|---|---|
| Molecule | Chains | Sequence Length | Organism | Details | Image |
| Cyanobacterial chloride importer | 232 | Mastigocladopsis repens | Mutation(s): 0 Membrane Entity: Yes | 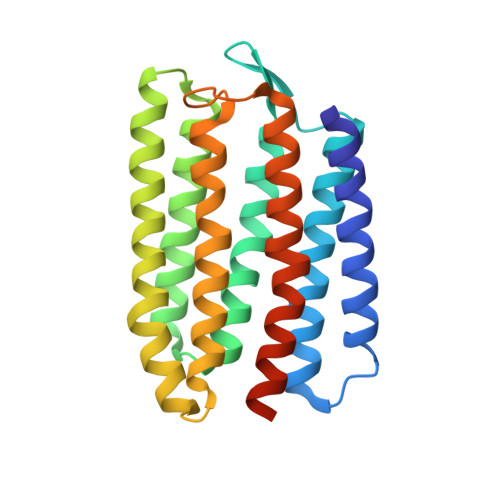 | |
UniProt | |||||
Find proteins for A0A6P3CW41 (Mastigocladopsis repens) Explore A0A6P3CW41 Go to UniProtKB: A0A6P3CW41 | |||||
Entity Groups | |||||
| Sequence Clusters | 30% Identity50% Identity70% Identity90% Identity95% Identity100% Identity | ||||
| UniProt Group | A0A6P3CW41 | ||||
Sequence AnnotationsExpand | |||||
Reference Sequence | |||||
| Ligands 3 Unique | |||||
|---|---|---|---|---|---|
| ID | Chains | Name / Formula / InChI Key | 2D Diagram | 3D Interactions | |
| RET Download:Ideal Coordinates CCD File | B [auth A] | RETINAL C20 H28 O NCYCYZXNIZJOKI-OVSJKPMPSA-N |  | ||
| OLA (Subject of Investigation/LOI) Download:Ideal Coordinates CCD File | C [auth A] D [auth A] E [auth A] F [auth A] G [auth A] | OLEIC ACID C18 H34 O2 ZQPPMHVWECSIRJ-KTKRTIGZSA-N |  | ||
| BR Download:Ideal Coordinates CCD File | Q [auth A], R [auth A] | BROMIDE ION Br CPELXLSAUQHCOX-UHFFFAOYSA-M |  | ||
| Length ( Å ) | Angle ( ˚ ) |
|---|---|
| a = 61.618 | α = 90 |
| b = 61.618 | β = 90 |
| c = 240.176 | γ = 120 |
| Software Name | Purpose |
|---|---|
| BUSTER | refinement |
| MxDC | data collection |
| HKL-2000 | data scaling |
| PHENIX | phasing |
| Coot | model building |
| Funding Organization | Location | Grant Number |
|---|---|---|
| National Research Foundation (Korea) | Korea, Republic Of | 20171A2B2008483 |
| National Research Foundation (Korea) | Korea, Republic Of | 20161A6A3A04010213 |