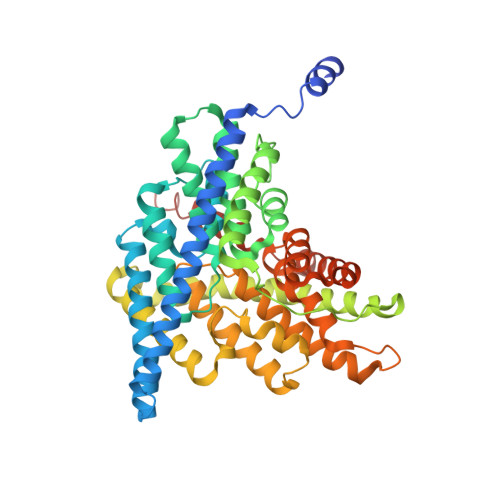
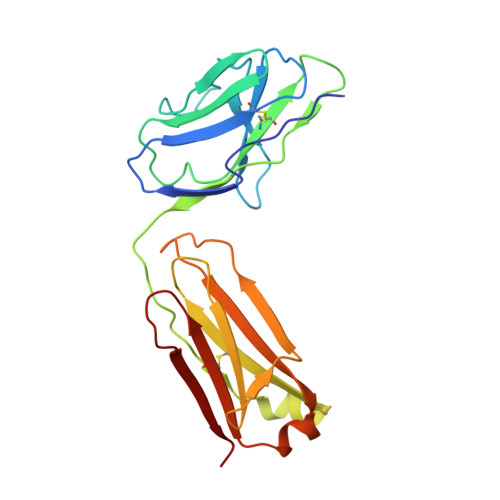
Mutation of external glutamate residue reveals a new intermediate transport state and anion binding site in a CLC Cl-/H+antiporter.
Park, K., Lee, B.C., Lim, H.H.(2019) Proc Natl Acad Sci U S A 116: 17345-17354
- PubMed: 31409705 Search on PubMedSearch on PubMed Central
- DOI: https://doi.org/10.1073/pnas.1901822116
- Primary Citation Related Structures:
6AD7, 6AD8, 6ADA, 6ADB, 6ADC, 6K5A, 6K5D, 6K5F, 6K5I - PubMed Abstract:
The CLC family of proteins are involved in a variety of physiological processes to control cellular chloride concentration. Two distinct classes of CLC proteins, Cl - channels and Cl - /H + antiporters, have been functionally and structurally investigated over the last several decades. Previous studies have suggested that the conformational heterogeneity of the critical glutamate residue, Glu ex , could explain the transport cycle of CLC-type Cl - /H + antiporters. However, the presence of multiple conformations ( Up , Middle , and Down ) of the Glu ex has been suggested from combined structural snapshots of 2 different CLC antiporters: CLC-ec1 from Escherichia coli and cmCLC from a thermophilic red alga, Cyanidioschyzon merolae Thus, we aimed to investigate further the heterogeneity of Glu ex -conformations in CLC-ec1, the most deeply studied CLC antiporter, at both functional and structural levels. Here, we show that the crystal structures of the Glu ex mutant E148D and wild-type CLC-ec1 with varying anion concentrations suggest a structural intermediate, the " Midlow " conformation. We also found that an extra anion can be located above the external Cl - -binding site in the E148D mutant when the anion concentration is high. Moreover, we observed that a carboxylate in solution can occupy either the external or central Cl - -binding site in the ungated E148A mutant using an anomalously detectable short carboxylic acid, bromoacetate. These results lend credibility to the idea that the Glu ex can take at least 3 distinct conformational states during the transport cycle of a single CLC antiporter.
- Department of Structure and Function of Neural Network, Korea Brain Research Institute, 41068 Daegu, Republic of Korea.
Organizational Affiliation: