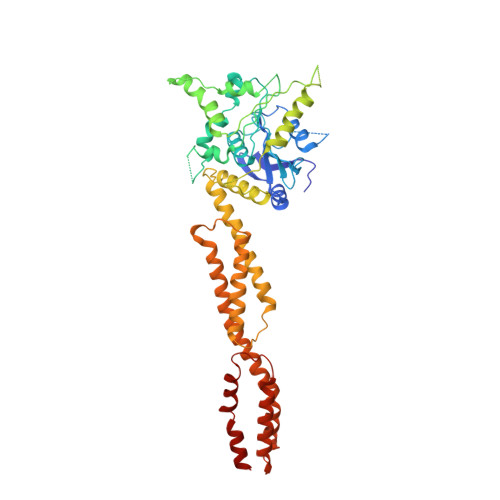
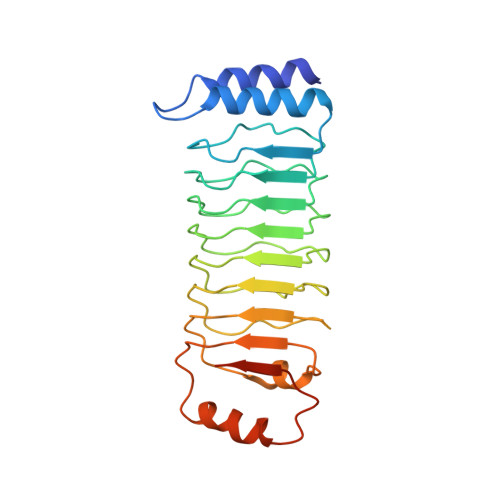
Structural mechanism for guanylate-binding proteins (GBPs) targeting by the Shigella E3 ligase IpaH9.8.
Ji, C., Du, S., Li, P., Zhu, Q., Yang, X., Long, C., Yu, J., Shao, F., Xiao, J.Y.(2019) PLoS Pathog 15: e1007876-e1007876
- PubMed: 31216343 Search on PubMedSearch on PubMed Central
- DOI: https://doi.org/10.1371/journal.ppat.1007876
- Primary Citation Related Structures:
6K1Z, 6K2D - PubMed Abstract:
The guanylate-binding proteins (GBPs) belong to the dynamin superfamily of GTPases and function in cell-autonomous defense against intracellular pathogens. IpaH9.8, an E3 ligase from the pathogenic bacterium Shigella flexneri, ubiquitinates a subset of GBPs and leads to their proteasomal degradation. Here we report the structure of a C-terminally truncated GBP1 in complex with the IpaH9.8 Leucine-rich repeat (LRR) domain. IpaH9.8LRR engages the GTPase domain of GBP1, and differences in the Switch II and α3 helix regions render some GBPs such as GBP3 and GBP7 resistant to IpaH9.8. Comparisons with other IpaH structures uncover interaction hot spots in their LRR domains. The C-terminal region of GBP1 undergoes a large rotation compared to previously determined structures. We further show that the C-terminal farnesylation modification also plays a role in regulating GBP1 conformation. Our results suggest a general mechanism by which the IpaH proteins target their cellular substrates and shed light on the structural dynamics of the GBPs.
- The State Key Laboratory of Protein and Plant Gene Research, School of Life Sciences, Peking-Tsinghua Center for Life Sciences, Peking University, Beijing, China.
Organizational Affiliation: