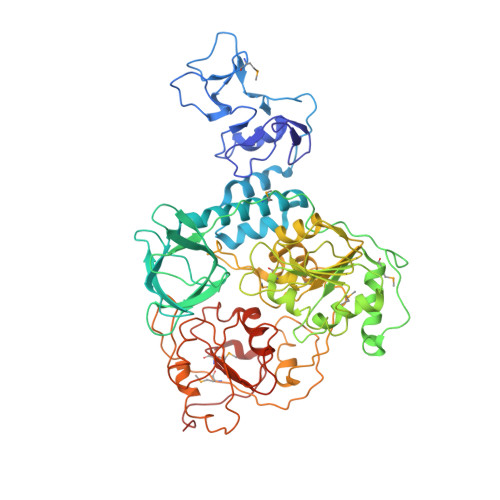
Delicate structural coordination of the Severe Acute Respiratory Syndrome coronavirus Nsp13 upon ATP hydrolysis.
Jia, Z., Yan, L., Ren, Z., Wu, L., Wang, J., Guo, J., Zheng, L., Ming, Z., Zhang, L., Lou, Z., Rao, Z.(2019) Nucleic Acids Res 47: 6538-6550
- PubMed: 31131400 Search on PubMedSearch on PubMed Central
- DOI: https://doi.org/10.1093/nar/gkz409
- Primary Citation Related Structures:
6JYT - PubMed Abstract:
To date, an effective therapeutic treatment that confers strong attenuation toward coronaviruses (CoVs) remains elusive. Of all the potential drug targets, the helicase of CoVs is considered to be one of the most important. Here, we first present the structure of the full-length Nsp13 helicase of SARS-CoV (SARS-Nsp13) and investigate the structural coordination of its five domains and how these contribute to its translocation and unwinding activity. A translocation model is proposed for the Upf1-like helicase members according to three different structural conditions in solution characterized through H/D exchange assay, including substrate state (SARS-Nsp13-dsDNA bound with AMPPNP), transition state (bound with ADP-AlF4-) and product state (bound with ADP). We observed that the β19-β20 loop on the 1A domain is involved in unwinding process directly. Furthermore, we have shown that the RNA dependent RNA polymerase (RdRp), SARS-Nsp12, can enhance the helicase activity of SARS-Nsp13 through interacting with it directly. The interacting regions were identified and can be considered common across CoVs, which provides new insights into the Replication and Transcription Complex (RTC) of CoVs.
- Laboratory of Structural Biology, School of Medicine, Tsinghua University, Beijing 100084, China.
Organizational Affiliation: