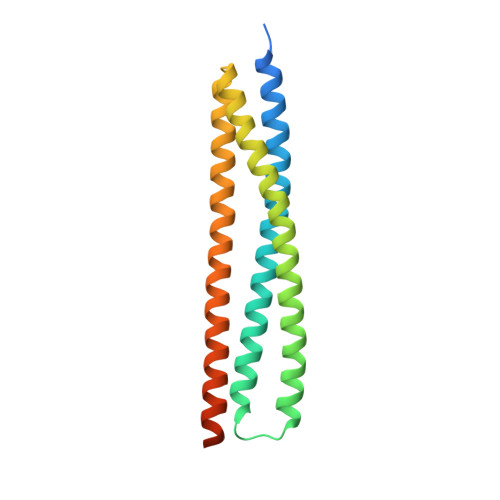
Structural insights into a HECT-type E3 ligase AREL1 and its ubiquitination activitiesin vitro.
Singh, S., Ng, J., Nayak, D., Sivaraman, J.(2019) J Biological Chem 294: 19934-19949
- PubMed: 31732561 Search on PubMedSearch on PubMed Central
- DOI: https://doi.org/10.1074/jbc.RA119.010327
- Primary Citation Related Structures:
6JX5, 6JX6 - PubMed Abstract:
The HECT E3 ligase family comprises three subfamilies: NEDD4 E3 ubiquitin protein ligase (NEDD4), HECT and RLD domain-containing E3 ubiquitin protein ligase (HERC), and "other." Most previous studies have focused on the NEDD4 subfamily. Apoptosis-resistant E3 ligase 1 (AREL1) belongs to "other" subfamily HECT that inhibits apoptosis by ubiquitinating and degrading proapoptotic proteins. Here, we report the crystal structure of the extended HECT domain of AREL1 (amino acids (aa) 436-823) at 2.4 Å resolution and its ubiquitination of the proapoptotic protein second mitochondria-derived activator of caspase (SMAC). We found that the extended HECT domain adopts an inverted, T-shaped, bilobed conformation and harbors an additional loop (aa 567-573) absent in all other HECT members. We also show that the N-terminal extended region (aa 436-482) preceding the HECT domain is indispensable for its stability and activity and that without this region, the HECT domain becomes inactive. AREL1 ubiquitinated SMAC, primarily on Lys 62 and Lys 191 We solved the crystal structure of the tetrameric form of SMAC to 2.8 Å resolution, revealing the Lys 62 and Lys 191 locations. The AREL1 HECT domain assembled Lys 33 -, Lys 48 -, and Lys 63 -linked polyubiquitin chains. Moreover, E701A substitution in the AREL1 HECT domain substantially increased its autopolyubiquitination and SMAC ubiquitination activity, whereas deletion of the last three amino acids at the C terminus completely abrogated AREL1 autoubiquitination and reduced SMAC ubiquitination. Finally, an AREL1-specific ubiquitin variant inhibited SMAC ubiquitination in vitro Our findings may assist in the development of AREL1 inhibitors that block its anti-apoptotic activity in cancer.
- Department of Biological Sciences, 14 Science Drive 4, National University of Singapore, 117543 Singapore.
Organizational Affiliation: