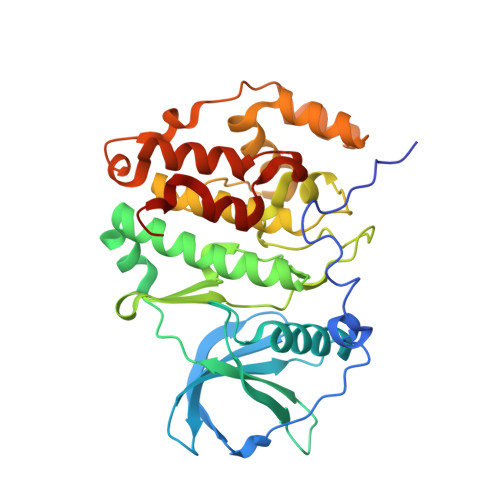
A promiscuous kinase inhibitor delineates the conspicuous structural features of protein kinase CK2a1.
Tsuyuguchi, M., Nakaniwa, T., Sawa, M., Nakanishi, I., Kinoshita, T.(2019) Acta Crystallogr F Struct Biol Commun 75: 515-519
- PubMed: 31282872 Search on PubMedSearch on PubMed Central
- DOI: https://doi.org/10.1107/S2053230X19008951
- Primary Citation Related Structures:
6JWA - PubMed Abstract:
Protein kinase CK2a1 is a serine/threonine kinase that plays a crucial role in the growth, proliferation and survival of cells and is a well known target for tumour and glomerulonephritis therapies. Here, the crystal structure of the kinase domain of CK2a1 complexed with 5-iodotubercidin (5IOD), an ATP-mimetic inhibitor, was determined at 1.78 Å resolution. The structure shows distinct structural features and, in combination with a comparison of the crystal structures of five off-target kinases complexed with 5IOD, provides valuable information for the development of highly selective inhibitors.
- Graduate School of Science, Osaka Prefecture University, 1-1 Gakuen-cho, Naka-ku, Sakai, Osaka 599-8531, Japan.
Organizational Affiliation: