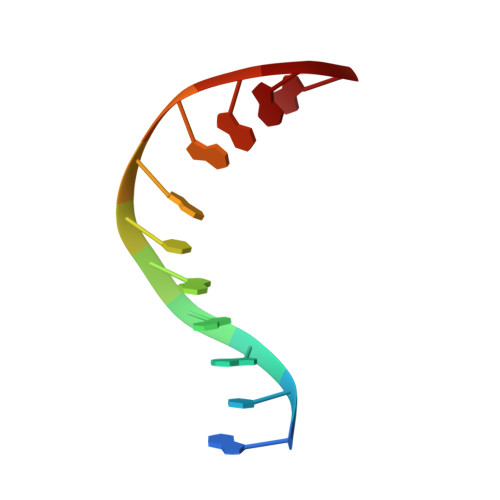
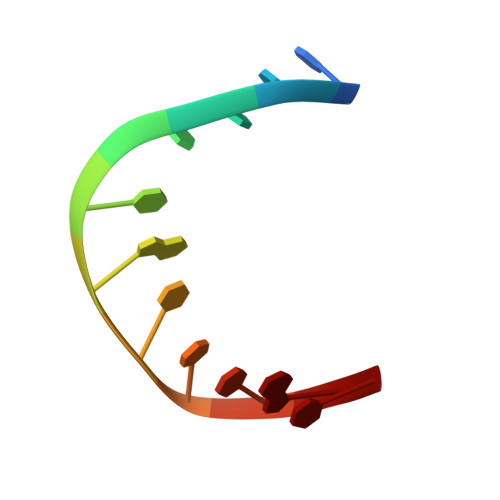
The crystal structure of Capicua HMG-box domain complexed with the ETV5-DNA and its implications for Capicua-mediated cancers.
Lee, H., Song, J.J.(2019) FEBS J 286: 4951-4963
- PubMed: 31323153 Search on PubMed
- DOI: https://doi.org/10.1111/febs.15008
- Primary Citation Related Structures:
6JRP - PubMed Abstract:
Capicua (CIC) is a transcriptional repressor and functions downstream of the receptor tyrosine kinase (RTK) signaling pathway. Somatic mutations found in the HMG-box DNA binding domain in CIC have been implicated in several cancers such as oligodendroglioma, oligoastrocytoma, and adenocarcinoma. However, the molecular basis of the DNA binding of CIC and the effect of the somatic mutations found in cancers on DNA binding have not been investigated. Here, we report the crystal structure of the HMG-box domain of CIC complexed with its target DNA, the promoter of Ets Translocation Variant 5 (ETV5). The structure shows that the HMG-box domain has an L-shaped structure and recognizes the minor groove leading to DNA bending. Our structure combined with an electrophoretic mobility shift assay (EMSA) revealed that cancer-associated mutations in the HMG-box domain abrogate the interaction with DNA. These results provide the molecular insight into the DNA binding of CIC and reveal the effects of carcinogenic mutations on DNA binding.
- Department of Biological Sciences, Korea Advanced Institute of Science and Technology (KAIST), Daejeon, Korea.
Organizational Affiliation: