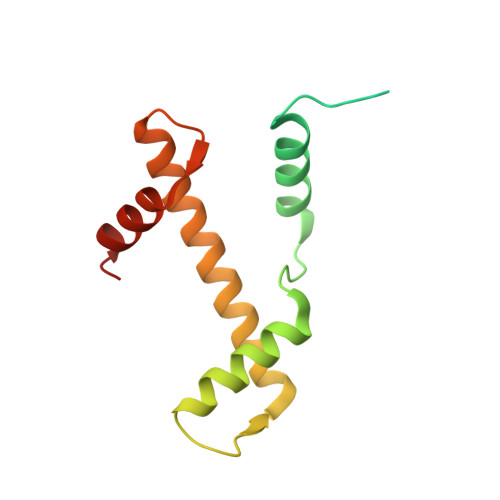
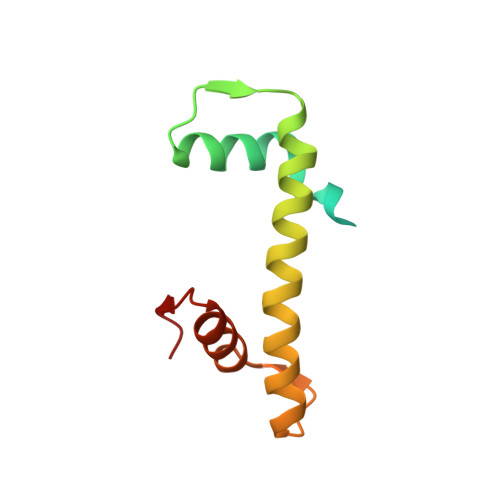
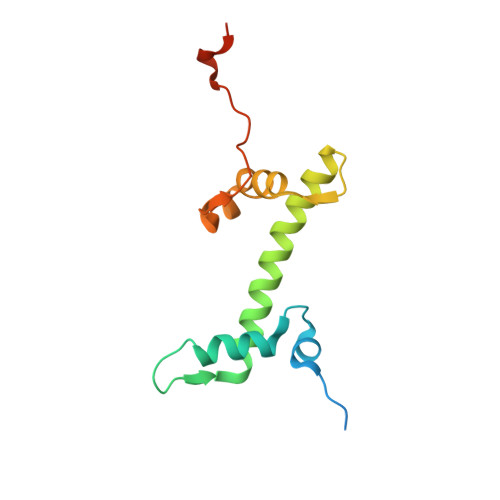
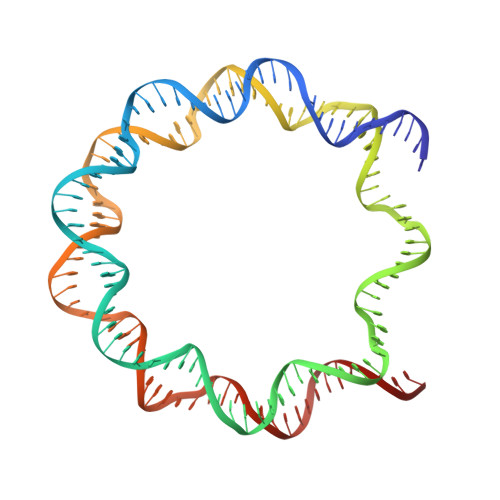
Structure-based design of an H2A.Z.1 mutant stabilizing a nucleosome in vitro and in vivo.
Horikoshi, N., Kujirai, T., Sato, K., Kimura, H., Kurumizaka, H.(2019) Biochem Biophys Res Commun 515: 719-724
- PubMed: 31186139 Search on PubMed
- DOI: https://doi.org/10.1016/j.bbrc.2019.06.012
- Primary Citation Related Structures:
6JOU - PubMed Abstract:
The nucleosome containing the histone H2A.Z.1 variant is unstable, as compared to the canonical nucleosome in vitro, and the incorporation of H2A.Z.1 into chromatin is less stable than that of the canonical H2A in vivo. In the present study, we designed a human H2A.Z.1(S42R) mutant, in which the Ser42 residue is replaced by Arg. In the crystal structure of the nucleosome containing H2A.Z.1(S42R), the Arg residue inserted at the H2A.Z.1-Ser42 position forms additional hydrogen bonds and electrostatic interactions with the DNA backbone phosphates. The Arg42 residue is located in the L1-loop region of H2A.Z.1, but the backbone geometry of the L1-loop is not affected by the H2A.Z.1(S42R) substitution. The nucleosome containing H2A.Z.1(S42R) exhibited enhanced thermal stability, as compared to that containing wild-type H2A.Z.1 in vitro. Fluorescence recovery after photobleaching experiments revealed that H2A.Z.1(S42R) was more stably incorporated in chromatin than wild-type H2A.Z.1 in living cells. Therefore, the H2A.Z.1(S42R) mutant stabilizes the nucleosome in vitro and in vivo, and may be useful as a tool to study the functional significance of the unstable nature of the H2A.Z.1 nucleosome.
- Laboratory of Structural Biology, Graduate School of Advanced Science and Engineering, Waseda University, 2-2 Wakamatsu-cho, Shinjuku-ku, Tokyo, 162-8480, Japan; Life Science Center for Survival Dynamics, Tsukuba Advanced Research Alliance (TARA), University of Tsukuba, 1-1-1 Tennodai, Tsukuba, Ibaraki, 305-8575, Japan.
Organizational Affiliation: