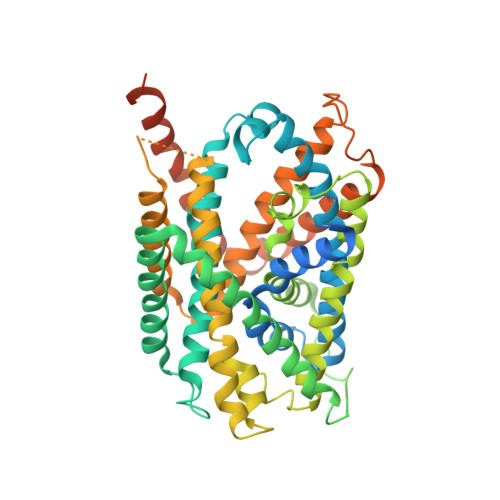
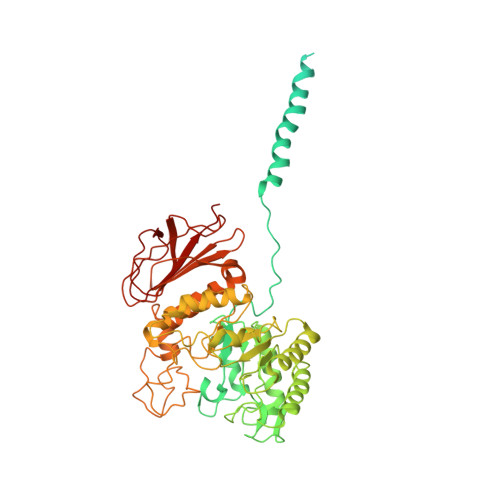
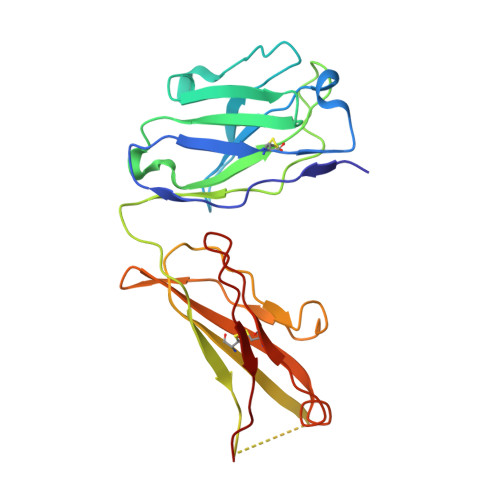
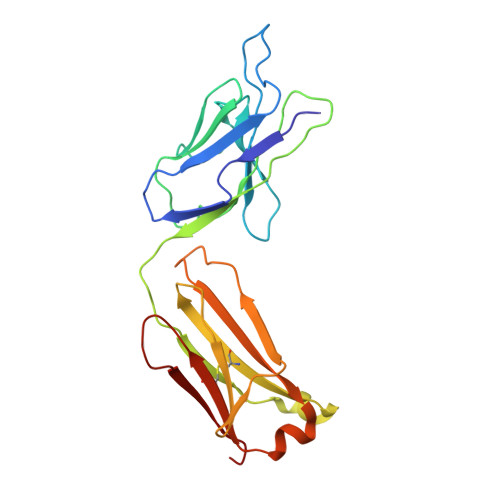
Cryo-EM structure of the human L-type amino acid transporter 1 in complex with glycoprotein CD98hc.
Lee, Y., Wiriyasermkul, P., Jin, C., Quan, L., Ohgaki, R., Okuda, S., Kusakizako, T., Nishizawa, T., Oda, K., Ishitani, R., Yokoyama, T., Nakane, T., Shirouzu, M., Endou, H., Nagamori, S., Kanai, Y., Nureki, O.(2019) Nat Struct Mol Biol 26: 510-517
- PubMed: 31160781 Search on PubMed
- DOI: https://doi.org/10.1038/s41594-019-0237-7
- Primary Citation Related Structures:
6JMQ, 6JMR - PubMed Abstract:
The L-type amino acid transporter 1 (LAT1 or SLC7A5) transports large neutral amino acids across the membrane and is crucial for brain drug delivery and tumor growth. LAT1 forms a disulfide-linked heterodimer with CD98 heavy chain (CD98hc, 4F2hc or SLC3A2), but the mechanism of assembly and amino acid transport are poorly understood. Here we report the cryo-EM structure of the human LAT1-CD98hc heterodimer at 3.3-Å resolution. LAT1 features a canonical Leu T-fold and exhibits an unusual loop structure on transmembrane helix 6, creating an extended cavity that might accommodate bulky amino acids and drugs. CD98hc engages with LAT1 through the extracellular, transmembrane and putative cholesterol-mediated interactions. We also show that two anti-CD98 antibodies recognize distinct, multiple epitopes on CD98hc but not its glycans, explaining their robust reactivities. These results reveal the principles of glycoprotein-solute carrier assembly and provide templates for improving preclinical drugs and antibodies targeting LAT1 or CD98hc.
- Department of Biological Sciences, Graduate School of Science, The University of Tokyo, Tokyo, Japan.
Organizational Affiliation: