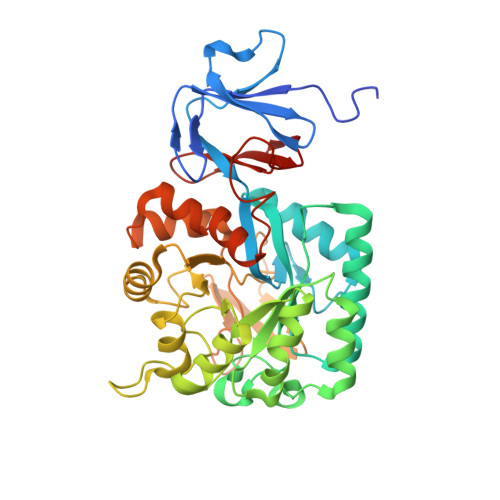
Quaternary variations in the structural assembly of N-acetylglucosamine-6-phosphate deacetylase from Pasteurella multocida.
Manjunath, L., Coombes, D., Davies, J., Dhurandhar, M., Tiwari, V.R., Dobson, R.C.J., Sowdhamini, R., Ramaswamy, S., Bose, S.(2020) Proteins
- PubMed: 32865821 Search on PubMed
- DOI: https://doi.org/10.1002/prot.25996
- Primary Citation Related Structures:
6JKU - PubMed Abstract:
N-acetylglucosamine 6-phosphate deacetylase (NagA) catalyzes the conversion of N-acetylglucosamine-6-phosphate to glucosamine-6-phosphate in amino sugar catabolism. This conversion is an essential step in the catabolism of sialic acid in several pathogenic bacteria, including Pasteurella multocida, and thus NagA is identified as a potential drug target. Here, we report the unique structural features of NagA from P. multocida (PmNagA) resolved to 1.95 Å. PmNagA displays an altered quaternary architecture with unique interface interactions compared to its close homolog, the Escherichia coli NagA (EcNagA). We confirmed that the altered quaternary structure is not a crystallographic artifact using single particle electron cryo-microscopy. Analysis of the determined crystal structure reveals a set of hot-spot residues involved in novel interactions at the dimer-dimer interface. PmNagA binds to one Zn 2+ ion in the active site and demonstrates kinetic parameters comparable to other bacterial homologs. Kinetic studies reveal that at high substrate concentrations (~10-fold the K M ), the tetrameric PmNagA displays hysteresis similar to its distant neighbor, the dimeric Staphylococcus aureus NagA (SaNagA). Our findings provide key information on structural and functional properties of NagA in P. multocida that could be utilized to design novel antibacterials.
- Institute for Stem Cell Science and Regenerative Medicine, NCBS, GKVK Campus, Bangalore, Karnataka, India.
Organizational Affiliation: