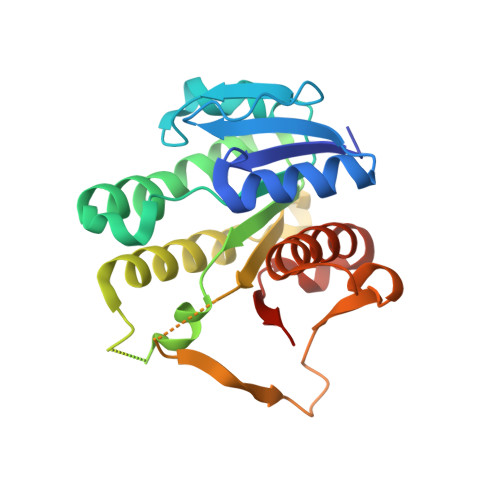
Crystal structures of human NSDHL and development of its novel inhibitor with the potential to suppress EGFR activity.
Kim, D.G., Cho, S., Lee, K.Y., Cheon, S.H., Yoon, H.J., Lee, J.Y., Kim, D., Shin, K.S., Koh, C.H., Koo, J.S., Choi, Y., Lee, H.H., Oh, Y.K., Jeong, Y.S., Chung, S.J., Baek, M., Jung, K.Y., Lim, H.J., Kim, H.S., Park, S.J., Lee, J.Y., Lee, S.J., Lee, B.J.(2021) Cell Mol Life Sci 78: 207-225
- PubMed: 32140747 Search on PubMedSearch on PubMed Central
- DOI: https://doi.org/10.1007/s00018-020-03490-2
- Primary Citation Related Structures:
6JKG, 6JKH - PubMed Abstract:
NAD(P)-dependent steroid dehydrogenase-like (NSDHL), an essential enzyme in human cholesterol synthesis and a regulator of epidermal growth factor receptor (EGFR) trafficking pathways, has attracted interest as a therapeutic target due to its crucial relevance to cholesterol-related diseases and carcinomas. However, the development of pharmacological agents for targeting NSDHL has been hindered by the absence of the atomic details of NSDHL. In this study, we reported two X-ray crystal structures of human NSDHL, which revealed a detailed description of the coenzyme-binding site and the unique conformational change upon the binding of a coenzyme. A structure-based virtual screening and biochemical evaluation were performed and identified a novel inhibitor for NSDHL harboring suppressive activity towards EGFR. In EGFR-driven human cancer cells, treatment with the potent NSDHL inhibitor enhanced the antitumor effect of an EGFR kinase inhibitor. Overall, these findings could serve as good platforms for the development of therapeutic agents against NSDHL-related diseases.
- Research Institute of Pharmaceutical Sciences, College of Pharmacy, Seoul National University, Seoul, 08826, Republic of Korea.
Organizational Affiliation: