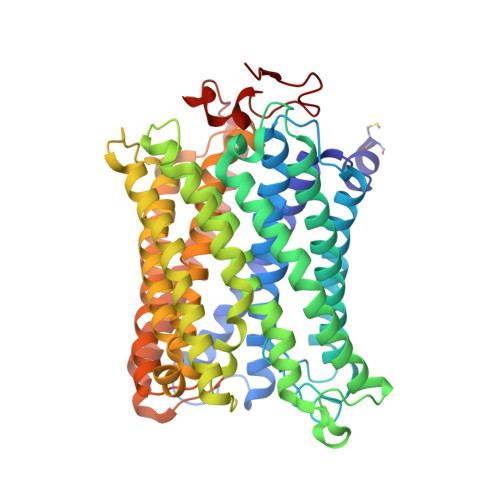
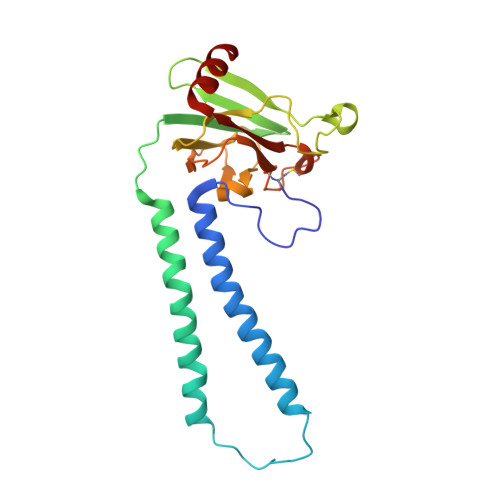
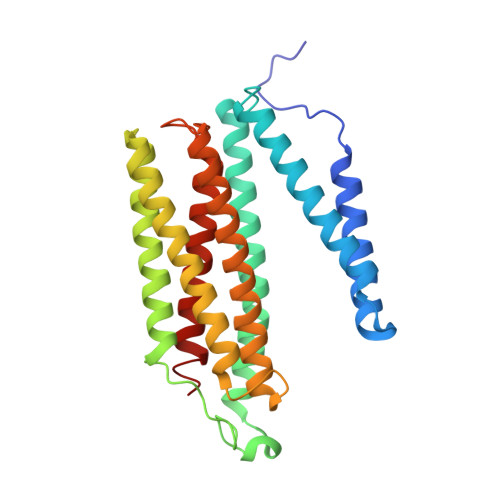
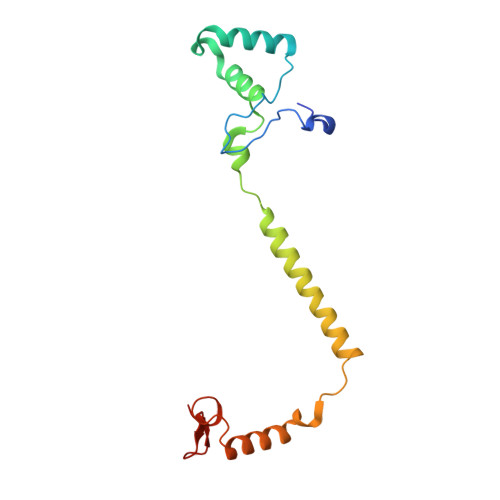
Low-dose X-ray structure analysis of cytochrome c oxidase utilizing high-energy X-rays.
Ueno, G., Shimada, A., Yamashita, E., Hasegawa, K., Kumasaka, T., Shinzawa-Itoh, K., Yoshikawa, S., Tsukihara, T., Yamamoto, M.(2019) J Synchrotron Radiat 26: 912-921
- PubMed: 31274413 Search on PubMedSearch on PubMed Central
- DOI: https://doi.org/10.1107/S1600577519006805
- Primary Citation Related Structures:
6J8M - PubMed Abstract:
To investigate the effect of high-energy X-rays on site-specific radiation-damage, low-dose diffraction data were collected from radiation-sensitive crystals of the metal enzyme cytochrome c oxidase. Data were collected at the Structural Biology I beamline (BL41XU) at SPring-8, using 30 keV X-rays and a highly sensitive pixel array detector equipped with a cadmium telluride sensor. The experimental setup of continuous sample translation using multiple crystals allowed the average diffraction weighted dose per data set to be reduced to 58 kGy, and the resulting data revealed a ligand structure featuring an identical bond length to that in the damage-free structure determined using an X-ray free-electron laser. However, precise analysis of the residual density around the ligand structure refined with the synchrotron data showed the possibility of a small level of specific damage, which might have resulted from the accumulated dose of 58 kGy per data set. Further investigation of the photon-energy dependence of specific damage, as assessed by variations in UV-vis absorption spectra, was conducted using an on-line spectrometer at various energies ranging from 10 to 30 keV. No evidence was found for specific radiation damage being energy dependent.
- SR Life Science Instrumentation Team, Life Science Research Infrastructure Group, Advanced Photon Technology Division, RIKEN SPring-8 Center, 1-1-1 Kouto, Sayo-cho, Sayo-gun, Hyogo 679-5148, Japan.
Organizational Affiliation: